Within our cohort, one patient was diagnosed with NS at birth and developed CML in blast phase (CML-BP) at 1.4 years of age (Table 1). Clinical features included short stature, motor delay, and valvular and supravalvular pulmonary stenosis, with a germline SOS1 R552S variant identified as the underlying genetic cause. No mutations were detected in JMML-associated genes. At presentation, the patient had severe anemia (hemoglobin 6.1 g/dl) and lymphoid blasts in blood. Bone marrow morphology revealed 30–40% lymphoid blasts in a markedly hypocellular marrow. Detection of a BCR::ABL1 fusion with a major breakpoint and the presence of BCR::ABL1 in 42% of myeloid interphase nuclei after cytoreductive therapy, confirmed the diagnosis of CML-BP. Treatment included reduced induction chemotherapy (two doses of vincristine and prednisolone) and tyrosine kinase inhibitor treatment. Due to refractory chylothorax, dasatinib was replaced by ponatinib after 15 months. On ponatinib, the patient achieved a major molecular response (BCR::ABL1/ABL1 ratio ≤1%), but no deeper molecular remission. Allogeneic hematopoietic stem cell transplantation was performed 3.3 years after diagnosis. Two years post-transplantation, the patient remains BCR::ABL1-negative despite subsequent graft failure.
Table 1 Germline variants in RAS signaling pathway in pediatric patients from CMLpaed II registry (reference genome GRCh37). The reference population was obtained from gnomAD™ v2.1.1 non-cancer (1).
The R552S variant resides in a strictly conserved codon and disrupts the autoinhibitory function of the protein through the lack of interaction with the side chains of D140 and D169 in the histone domain (Fig. 1A). SOS1 R552S was classified as pathogenic by ACMG and in silico predictors (mean score 0.823). In addition, the OncoVI tool for oncogenicity classification [11] labeled the variant as likely oncogenic. R552S belongs to a spectrum of known SOS1 variants in individuals with NS, with mean scores ranging from 0.086 to 0.969 (median 0.753; Supplementary Table 1). Within the specific context of NS-associated leukemia, R552S scored similarly to previously identified variants in pediatric patients with NS and acquired leukemia (M269R and M269T with a score of 0.898 and 0.871, respectively). While the SOS1 variants in CML are located in similar domains to those in acute leukemia, no cluster between the two cohorts can be defined (Fig. 1B).
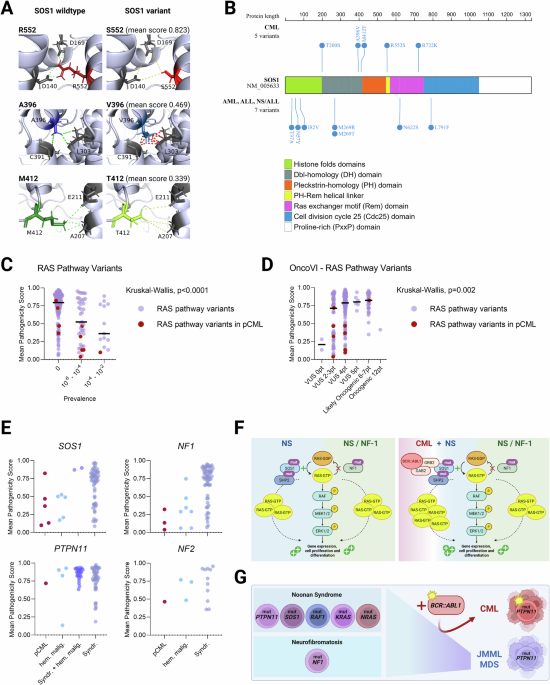
Fig. 1: Characteristics of germline variants in RAS pathway genes identified in pediatric patients with chronic myeloid leukemia (pCML).The alternative text for this image may have been generated using AI.
A Comparison of structural differences between the SOS1 wildtype and variants encoding the amino acid substitutions in the pCML cohort. Dotted lines represent the distance between amino acids in wildtype SOS1 (green), the modified distance (yellow), or possible clashes (red) in SOS1 variants. Protein structure and amino acid interactions illustrated with PyMOL software. B Distribution of germline missense SOS1 variants in pCML (upper part) and variants in individuals with acute leukemia or Noonan syndrome (NS)-associated leukemia (lower part). C Distribution of mean scores of all germline variants (SOS1, PTPN11, NF1, and NF2) identified in RASopathies and hematological malignancies and their prevalence in the healthy population, as provided by the Genome Aggregation Database (gnomAD™). D Distribution of mean scores of all germline variants across the OncoVI categories. The horizontal black lines indicate the median in the respective group. E Distribution of mean scores of germline variants across pCML, hematological malignancies (HM), syndromic hematological malignancies (Syndr. HM), and syndromic cases (Syndr.). F RAS/RAF/MAPK pathway in NS and neurofibromatosis type 1 (left). In the presence of the constitutively active BCR::ABL1 kinase, hyperactivated RAS leads to an additional increase in downstream processes (right). G Germline variants in the RAS signaling pathway are causative of NS and neurofibromatosis type 1. These individuals are predisposed to hematological disorders such as juvenile myelomonocytic leukemia (JMML) and myelodysplastic syndrome (MDS). The additional BCR::ABL1 translocation may trigger the early manifestation of CML. Created with BioRender.com.
Of the 145 pediatric patients for whom genetic analysis was available, 88 had a genetic variant in addition to the BCR::ABL1 fusion gene. A total of 10 patients (7%; five female and five male) were identified with 10 germline missense variants in four genes associated with the RAS/RAF/MAPK pathway, specifically SOS1, PTPN11, NF1, and NF2. The median age at diagnosis was 9.5 (1.4–16.8) years. Germline variants were discovered in six chronic phase CML, three de novo CML-BP, and one secondary CML-BP. According to ACMG guidelines, 7/10 variants were classified as variants of uncertain significance (VUS), 2/10 as likely benign, and 1/10 as pathogenic. Similarly, OncoVI classified 9/10 variants as VUS and 1/10 as likely oncogenic, corresponding to the pathogenic variant classified by ACMG. Their prevalence in the population database (0.0004–0.01%) indicates that they are not frequently occurring single-nucleotide polymorphisms.
When RAS pathway germline variants, both from pediatric patients with CML (n = 10) and those identified in previously published studies (n = 215), are stratified according to their prevalence in the reference population (gnomAD™ version 2.1.1, non-cancer) and their mean score is calculated, we observe that most variants (171/225; 76%) are not present in this database (Fig. 1C). There is a continuum within the CML variants from extremely rare variants predicted to have a more functional impact to the rather benign variants with higher prevalence. OncoVI classified 33/215 variants as likely oncogenic and 1/215 as oncogenic. The median of the predicted mean scores generated by the in silico tools increases with increasing assigned points by OncoVI (Fig. 1D). When we compare the groups separately in each gene, we observe that the mean scores attributed to the CML variants are similar to those of variants identified in hematological malignancies and in syndromic hematological disorders (Fig. 1E and Supplementary Fig. 1). There is significant variation in the median between the analyzed groups for the SOS1 (p = 0.01) and NF1 (p = 0.0006) genes. Variants received varying scores across the four prediction tools. Three in silico tools demonstrate the strongest correlation (r ≥ 0.804), whereas one exhibits a moderate correlation with the other tools (Supplementary Fig. 2).