Mice
All animal experiments were performed in accordance with institutional guidelines and were approved by the Institutional Animal Care and Use Committee (IACUC) of the University of Missouri (protocol number 65384). C57BL/6NCrl mice (Mus musculus) were obtained from Charles River Laboratories. Dpp4 flox/flox mice were generated by breeding targeted C57Bl/6NTac-DPP4tm1a Wtsi/Ics mice (European Mouse Mutant Cell Repository, EUCOMM) with 129S4/Bl6-Gt (ROSA) 26Sortm2(FLP*) Sor/J (stock no. 012930, The Jackson Laboratory). The offspring were further crossed with Vav-iCre mice (stock no. 018968, The Jackson Laboratory)70, to generate Dpp4fl/fl;Vav-Cre mice13. N-cad-tdTomato (Cdh2-tdTamato), N-cad-CreER (Cdh2-CreEr) and Gpc3 fl/fl strains were generated by L.L. at Stowers Institute for Medical Research16,18. Cxcl12fl/fl (stock no. 022457), Scf fl/fl (stock no. 017861) and Nestin-CreER (stock no. 016261) mice were purchased from The Jackson Laboratory. To induce expression of Cre-ER recombinase, mice received tamoxifen via intraperitoneal injection (Sigma, 75 mg tamoxifen per kilogram body weight) as described71. All mouse strains used in this study had a C57BL/6 genetic background. Both male and female mice aged 6–10 weeks were used unless otherwise specified. Animals were randomly assigned to experimental groups based on genotyping results. Investigators were blinded to group allocation during data analysis but not during experimental procedures. Sample sizes for each experiment are detailed in figure legends. Mice were housed in a specific-pathogen-free facility under a 12-h light/12-h dark cycle at an ambient temperature of 20–24 °C and relative humidity of 40–60%, with ad libitum access to food and water.
AML transplantation
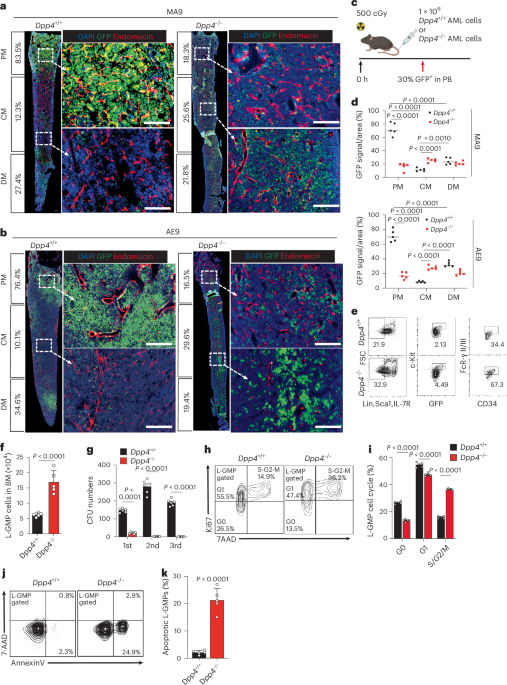
For Fig. 1a, we transplanted infected Dpp4-KO or control Lin− cells (2 × 105; infection efficiency consistently 40–50%) into lethally irradiated (1,000 cGy) C57/B6 mice (6–8 weeks old) and evaluated 2–50 weeks after transplantation. For other Dpp4 KO versus control AML models, 1 × 106 primary AML cells were transplanted C57/B6 recipients. For the homing assay, GFP+ AML cells were evaluated at 16 h. Engraftment was assessed 2 and 4 weeks after transplantation for MLL-AF9 and AML-ETO9a models, respectively. For control AML cells transplantation into N-cad+ mice with or without Cxcl12 or Gpc3, 500 L-GMPs were transplanted into sublethally irradiated (500 cGy) mice and engraftment assessed 2–8 weeks after transplantation. To preserve niche cell integrity, all mice were sublethally irradiated (500 cGy) before transplantation. The maximal leukaemia burden permitted under the approved IACUC protocol (#65384) was defined according to institutional humane endpoint criteria. Animals were monitored regularly and euthanized upon reaching predefined signs of morbidity or distress. The maximal leukaemia burden permitted by the ethics committee was not exceeded in any experiment.
Flow cytometry
PB, BM, spleen and liver haematopoietic cells were labelled with the following antibodies (all from BioLegend unless otherwise noted): anti-CD3e-PE/Cyanine5 (clone 17A2, #100310, 1:200), anti-Ly6G/Ly6C (Gr-1)-PE/Cyanine5 (clone RB6-8C5, #108410, 1:200), anti-CD11b-PE/Cyanine5 (clone M1/70, #101210, 1:200), anti-CD45R-PE/Cyanine5 (clone RA3-6B2, #103210, 1:200), anti-Ter-119-PE/Cyanine5 (clone TER-119, #116210, 1:200), anti-CD117 (c-Kit)-APC (clone 2B8, #105812, 1:200), anti-Sca-1-PE-Cy7 (clone D7, #108114, 1:200), anti-CD150-PE (clone TC15-12F12.2, #115904, 1:200), anti-CD48-APC/Cyanine7 (clone HM48-1, #103432, 1:200), anti-Ki67-FITC (clone 16A8, #652410, 1:200), Hoechst 34580 (BD Pharmingen, #565877), anti-CD16/32-PE (clone 93, #101308, 1:200), anti-CD34-FITC (clone RAM34, eBioscience, #11-0341-82, 1:200), anti-CD127-APC/Cyanine7 (clone A7R34, #135040, 1:200), anti-CD135-Brilliant Violet 421 (clone A2F10, #135314, 1:200), anti-CD45-APC (clone I3/2.3, #147708, 1:200), anti-Ter-119-PE (clone TER-119, #116208, 1:200), anti-CD31-PerCP/Cyanine5.5 (clone W18222B, #160206, 1:200), Annexin V (#640941, 1:20) and propidium iodide (#421301, 1:50). Intracellular staining was performed using the Foxp3/Transcription Factor Staining Kit (eBioscience) according to the manufacturer’s protocol. Flow cytometry analyses were performed independently in triplicate, with biological replicates from at least five mice per condition. Technical replicates were included for measurement accuracy.
Immunofluorescence staining and quantification
Femurs were perfused with phosphate-buffered saline (PBS), fixed with 4% paraformaldehyde and subjected to frozen sectioning. Antigen retrieval was performed with 1 μg ml−1 proteinase K in TE buffer (100 mM Tris–HCl, pH 8.0, 50 mM EDTA) at 37 °C for 30 min. Sections were blocked with Universal Blocking Reagent (BioGenex) and incubated overnight at 4 °C with primary antibodies, including anti-Endomucin (goat polyclonal, R&D Systems, #AF4666, 1:100), Biotin anti-mouse Lineage Panel (clone 145-2C11; RB6-8C5; RA3-6B2; Ter-119; M1/70 BioLegend, #133307, 1:200), Biotin anti-mouse IL-7Rα (clone A7R34, BioLegend, #135006, 1:200), Biotin anti-mouse Sca-1 (clone D7, BioLegend, #108104, 1:200), phycoerythrin (PE) anti-mouse CD34 (clone HM34, BioLegend, #128610, 1:200), Alexa Fluor 647-anti mouse CD117(c-kit) (clone 2B8, BioLegend, #105818, 1:200) and anti-GPC3 (rabbit polyclonal, Abcam, #ab216606, 1:200). Secondary antibodies included donkey anti-goat Alexa Fluor 555 (Invitrogen; 1:500), goat anti-rabbit Alexa Fluor 750 (Invitrogen; 1:500) and Brilliant Violet 421-conjugated streptavidin (BioLegend; 1:500) at room temperature for 1 h. A 4′,6-diamidino-2-phenylindole (DAPI) stock solution was diluted to 300 nM in PBS, and 300 ml was added to the coverslip preparation for 1 min. Sections were rinsed three times in PBS, excess buffer drained from the coverslip and mounted with Shandon Immu-Mount (Fisher Scientific). Image stitching was done to capture the entire specimen at high magnification and seamlessly create a single high-resolution image. Sections were imaged using a Keyence BZ-X800 fluorescence microscope at 20× magnification (resulting in 200× magnification) and 60× magnification (resulting in 600× magnification). Quantification was performed using Keyence BZ-X800 analyser software, assessing GFP+ AML cell distribution across BM areas (PM, CM and DM), L-GMP localization, and apoptosis. Distances between GFP+ AML cells and N-cad+ cells were measured using a minimum of 100 GFP+Kit+ and 80 GFP+Kit− AML cells per dataset. A minimum of five mice per condition were used for each quantification dataset. Detailed light microscopy acquisition parameters are provided in Supplementary Table 1.
Cytokine analyses
Cytokine quantification in plasma and BMEF were determined using the LEGENDplex Multi-Analyte Flow Assay Kit (BioLegend), a bead-based immunoassay that quantifies multiple cytokines simultaneously via flow cytometry. In brief, a custom mouse cytokine and chemokine panel was used to measure the concentrations of the designed cytokines/chemokines. The LSRFortessa X-20 Cell Analyzer (BD Biosciences) was used for data acquisition, and results were analysed using the LEGENDplex Data Analysis software. Assays were performed in 96-well plates following manufacturer protocols, with data recorded using a Fisherbrand microplate photometer. BMEF and plasma were used for enzyme-linked immunosorbent assay (ELISA) collected using the mouse SDF-1 alpha ELISA Kit (Invitrogen) following the manufacturer’s protocol.
Migration assay
In vitro, the Transwell migration assay was utilized to assess cell migration. DPP4+/+ and DPP4−/− AML cells were cultured and then seeded in serum-free medium into the upper chamber of Transwell inserts (Corning) with an 8-μm pore size, Matrigel-coated membrane. The upper wells contained CXCL12 at concentrations of 0 ng ml−1. The lower chambers were filled with culture medium containing 100 ng ml−1 CXCL12. After a 4-h incubation at 37 °C and 5% CO2, non-migratory cells on the upper membrane surface were removed using a cotton swab. Migratory cells on the lower membrane surface were visualized and quantified in multiple random fields under a microscope. Migration rates and statistical significance were analysed accordingly.
In vivo, mice (n = 5) were intravenously injected with PBS or CXCL12 (0–500 ng g−1 per mouse as indicated in Fig. 3k). The percentage of GFP+ AML cells in PB was measured before and 17 h after injection.
Colony assays
Mouse AML cells were diluted to the indicated concentration in Iscove’s modified Dulbecco’s medium with 2% fetal bovine serum and were then seeded into methylcellulose medium M3534 (STEMCELL Technologies) for myeloid colony formation analysis72. For serial CFU assays, L-GMP cells were re-isolated by fluorescence-activated cell sorting (FACS) from collected colonies and replated into fresh methylcellulose medium for subsequent rounds of colony formation. Each CFU assay was performed in at least triplicate using independent biological replicates derived from distinct AML mice. Consistent colony-forming capacity was observed across all replicates.
DPP4 activity assay
DPP4 activity was measured in plasma-EDTA, BMEF and cell lysates. For each assay, 20 μl of serum or BMEF, or 100 nM of GPC3 protein, was diluted in DPP4 assay buffer (Tris–HCl (pH 8.0), 150 mM NaCl and protease inhibitor cocktail) in a black 96-well plate to a final volume of 50 μl. An equal volume (50 μl) of 200 mM H-Ala-Pro-AFC substrate (I-1680; Bachem Americas) was added, and the plate was incubated for 10 min at room temperature in the dark. Fluorescence was measured using a Synergy Microplate Reader at excitation/emission wavelengths of 405/535 nm, and results were reported as relative light units (RLUs).
Ligand binding assay
Recombinant His-tagged GPC3 binding to Dpp4+/+ AML cells was assessed as similarly described73. In brief, 1 × 106 Dpp4+/+ AML cells were incubated with or without His-GPC3 (100 nM) in 200 ml PBS/1% BSA for 3 h at 25 °C. Non-specific binding was subtracted. After incubation, cells were washed twice by centrifugation, resuspended in ice-cold PBS/1% BSA, and stained with Alexa Fluor 488 anti-His Tag Antibody for flow cytometry analysis.
Quantitative RT–qPCR
Reverse-transcription quantitative PCR (RT–qPCR) was performed using 5 ng total RNA, gene-specific primers and a QIAGEN One Step RT-PCR kit (210210; Qiagen) following the manufacturer’s instructions. 18S rRNA was used as an internal control for normalization. The primer sequences used are listed below: TFPI: forward(5′-GGG CTC CGT TCT TGG TCT C-3′) and reverse(5′-TTG AAT CTG CGG CAC TTT TGC-3′), MMP9: forward(5′-CTG GAC AGC CAG ACA CTA AAG-3′) and reverse (5′-CTC GCG GCA AGT CTT CAG AG-3′), MMP2: forward(5′-CAA GTT CCC CGG CGA TGT C-3′) and reverse (5′-TTC TGG TCA AGG TCA CCT GTC-3′), MMP3: forward(5′-ACA TGG AGA CTT TGT CCC TTT TG-3′) and reverse (5′-TTG GCT GAG TGG TAG AGT CCC-3′), MMP13: forward(5′-CTT CTT CTT GTT GAG CTG GAC TC-3′) and reverse (5′-CTG TGG AGG TCA CTG TAG ACT-3′), MMP14: forward(5′-CAG TAT GGC TAC CTA CCT CCA G-3′) and reverse (5′-GCC TTG CCT GTC ACT TGT AAA-3′), cathepsin G: forward(5′-AGG GTT TCT GGT GCG AGA AG-3′) and reverse (5′-GTT CTG CGG ATT GTA ATC AGG AT-3′), Elastase: forward(5′-AGC AGT CCA TTG TGT GAA CGG-3′) and reverse (5′-CAC AGC CTC CTC GGA TGA AG-3′), DPP8: forward (5′-GGG AAA TGG TGA ATC ACA GGA C-3′) and reverse (5′-ATG TAG CCG TGG TAT TTT CTG G-3′), GPC3: forward (5′-CAG CCC GGA CTC AAA TGG G-3′) and reverse (5′-CAG CCG TGC TGT TAG TTG GTA-3′).
RNA-seq analysis
BM AML cells were FACS-sorted from two independent Dpp4+/+ and Dpp4−/− leukaemia-bearing mice, as well as from N-cad-Cre; Cxcl12+/+ and N-cad-Cre; Cxcl12−/− mice. Total RNA was extracted using the miRNeasy Mini Kit (QIAGEN) according to the manufacturer’s instructions. RNA concentration was quantified using the Qubit RNA HS Assay Kit (Invitrogen) on a Qubit 4 Fluorometer, and RNA integrity was assessed using the Fragment Analyzer automated capillary electrophoresis system (Agilent Technologies). For library preparation, 1 µg of total RNA was used. Polyadenylated mRNA was isolated, fragmented and reverse-transcribed to generate double-stranded cDNA. Adapter ligation and library amplification were performed using the TruSeq Stranded mRNA Library Prep Kit (Illumina) following the manufacturer’s protocol. Amplified libraries were purified using AxyPrep Mag PCR Clean-Up beads (Axygen). The final library quality and fragment size distribution were validated using the Fragment Analyzer, and concentrations were determined using the Qubit™ dsDNA HS Assay Kit (Invitrogen). Sequencing-ready libraries were diluted and pooled according to Illumina’s standard protocol and paired-end sequencing was performed on an Illumina NextSeq 500 platform at the University of Missouri DNA Core Facility.
Cell preparation for scRNA-seq
For tissue collection, femurs were collected after euthanasia and immediately placed in ice-cold PBS. The PM and DM regions of the bone were isolated. BM was extracted by crushing the bones with a mortar and pestle, followed by enzymatic digestion with collagenase/dispase at 37 °C for 45 min. The resulting cell suspension was washed, lysed and filtered through a 100-μm strainer. FACS isolation of non-haematopoietic cells: BM single-cell suspensions were stained with Anti-CD45-APC (clone I3/2.3, #147708, 1:200), anti-Ter-119-PE (clone TER-119, #116208, 1:200) and Anti-CD31-PerCP/Cyanine5.5 (clone W18222B, #160206, 1:200) to exclude haematopoietic cells and ECs. Live/dead discrimination was performed using DAPI. Viable CD45−Ter-119−CD31− (triple-negative) cells were sorted on a BD FACSAria II cell sorter.
Single-cell sequencing
Single cells were encapsulated into emulsion droplets using Chromium Controller (10x Genomics). scRNA-seq libraries were constructed using Chromium Single Cell 3′ v2 Reagent Kit according to the manufacturer’s protocol. In brief, the post-sorting sample volume was decreased, and cells were examined under a microscope and counted with a haemocytometer. Cells were then loaded in each channel with a target output of ~4,000 cells. Reverse transcription and library preparation were performed on C1000 Touch Thermal cycler with 96-Deep Well Reaction Module (Bio-Rad). Amplified cDNA and final libraries were evaluated on an Agilent Bioanalyzer using a High Sensitivity DNA Kit (Agilent Technologies). Individual libraries were diluted to 4 nM and pooled for sequencing. Pools were sequenced with 75 cycle run kits (26-bp Read1, 8-bp Index1 and 55-bp Read2) on the Novaseq 5000 Sequencing System (Illumina).
scRNA-seq analysis
Single-cell transcriptomic profiling was performed as previously described with modifications1. In brief, raw sequencing data were processed using the Cell Ranger Single Cell Software Suite (v3.0.2, 10x Genomics) for quality control, sample demultiplexing, barcode assignment and 3′ gene counting. Reads were aligned to the mouse reference transcriptome (refdata-cellranger-mm39-3.0.0) with default parameters. Gene–barcode matrices were subsequently imported into the Seurat v3 package (R software). Quality control filters were applied to exclude low-quality or apoptotic cells: cells with >5% mitochondrial unique molecular identifier content and <1,500 detected genes were removed. Genes expressed in fewer than two cells were also excluded. Following normalization and scaling, highly variable genes were identified and used for dimensionality reduction by PCA. Cell clustering was performed using the Seurat ‘FindNeighbors’ and ‘FindClusters’ functions with the first 20 principal components and a resolution parameter of 0.5. Uniform Manifold Approximation and Projection (UMAP) was applied for visualization in two-dimensional space. Differentially expressed genes (cluster-specific markers) were determined using the ‘FindMarkers/FindAllMarkers’ function with the Wilcoxon rank-sum test. To define cell identities, we compared cluster-specific gene signatures with canonical markers reported in prior literature1. For MSCs, markers including Lepr, Cdh2 (N-cadherin), Prx1, Osx (Sp7) and Nestin were examined. OLCs were identified by Bglap expression; fibroblasts by S100a4; chondrocytes by Acan and Col2a1; ECs by Cdh5; and pericytes by Acta2. Marker selection was based on well-established stromal lineage-defining studies and enabled robust annotation of BM niche subpopulations. All plots were generated in R using Seurat visualization functions.
Spleen histology
The spleen was fixed in 4% phosphate-buffered formalin, dehydrated, embedded in frozen section compound 22 blue (Leica), sectioned at 6 µm and placed onto coated Superfrost Plus Microscope Slides (Fisher Scientific). Sections were stained with haematoxylin and eosin using the Thermo Scientific Shandon Rapid-Chrome H&E Frozen Section Staining Kit. Images were acquired using a BZ-8000 fluorescence microscope with 20× and 60× objectives (yielding 200× and 600× magnifications).
Western blotting
Cells were lysed in Laemmli sample buffer (Sigma-Aldrich) supplemented with protease inhibitor cocktail (Roche Diagnostics). Samples were separated on SDS–PAGE gels (Bio-Rad) and transferred to nitrocellulose membranes (Bio-Rad) for protein detection. Primary antibodies were obtained from Cell Signaling Technology: Phospho-p44/42 MAPK (Erk1/2) (Thr202/Tyr204) (clone 20G11, Cell Signaling Technology, #75796S, 1:1,000), Phospho-NF-κB p65 (clone 93H1, Cell Signaling Technology, #3039S, 1:1,000), Phospho-STAT3 (Tyr705) (clone D3A7, Cell Signaling Technology, #9145S, 1:2,000), Phospho-p38 MAPK (clone D3F9, Cell Signaling Technology, #4092S, 1:1,000) and β-actin (clone C4, Santa Cruz Biotechnology, #sc-47778, 1:2,000). Secondary detection was performed with HRP-conjugated anti-rabbit and anti-mouse antibodies (R&D Systems, #HAF007 and #HAF005, respectively, 1:1,000), and protein bands were visualized using a chemiluminescent substrate (Invitrogen).
Complete blood count assay
PB was collected from mice by retro-orbital bleeding into EDTA-coated microtubes to prevent coagulation. Complete blood counts, including haemoglobin concentration, total leukocyte count and platelet count, were measured using an automated haematology analyser (Hemavet 950FS, Drew Scientific) according to the manufacturer’s instructions (performed by Comparative Clinical Pathology Services LLC). Each group contained five mice (n = 5), and results are presented as mean ± standard error of the mean (s.e.m.).
Statistics and reproducibility
Data are expressed as mean ± s.e.m. Statistical analyses were performed using GraphPad Prism Version 9.0 (GraphPad Prism Software). For continuous variables, normality and homogeneity of variance were assessed using the Shapiro–Wilk and Brown–Forsythe tests, respectively. After confirming homogeneous variances and normality, two-group comparisons for means were performed using the two-sided Student t-test, and multigroup comparisons for means were performed using two-way analysis of variance with Holm–Šidák multiple comparison test. For data that did not pass either normality or equal variance test, two-group comparisons were performed using the Mann–Whitney rank-sum test, and multigroup comparisons were performed using the Kruskal–Wallis one-way analysis of variance on ranks test with the Dunn post-hoc test. P < 0.05 was considered statistically significant. Animals were randomly assigned to experimental groups based on genotyping results using a simple randomization approach. Where applicable, littermates were distributed across treatment groups to minimize bias. Investigators were blinded to group allocation during data analysis but not during experimental procedures. No animals were excluded from the analyses. Data points were excluded only if predefined technical criteria were not met (for example, sample processing failure), before statistical analysis.
Sample size determination
No statistical methods were used to predetermine sample sizes. Sample sizes were selected on the basis of prior experience with the AML transplantation and BM niche models and are comparable to those reported in previous publications using similar experimental systems13,74. The chosen sample sizes are consistent with established standards in the field and were sufficient to detect biologically meaningful differences with appropriate statistical tests.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.