Sample acquisition, ethics and patient consent
Foetal liver samples were obtained from the MRC–Wellcome Trust – funded Human Developmental Biology Resource (HDBR; http://www.hdbr.org61) with appropriate maternal written consent and approval from the North East – Newcastle & North Tyneside 1 Research Ethics Committee. HDBR is regulated by the UK Human Tissue Authority (HTA; www.hta.gov.uk) and operates in accordance with the relevant HTA Codes of Practice.
Leukaemia samples from bone marrow aspirates or peripheral blood were collected through studies approved by the UK NHS research ethics committee. The tissue sources are: Great Ormond Street Hospital for Children diagnostic archives, as approved by a National Research Ethics Service Committee (London Brent, reference 16/LO/0960); and the AML Berlin-Frankfurt-Münster (BFM) study group. Bulk transcriptomic processed data with approval from the Goethe University Frankfurt (Ethics # 2021-341 and # 2023-1565). Ethical approval was obtained from the London – Brent Research Ethics Committee, the University Duisburg-Essen (Ethics #17-7462), Hannover Medical School (Ethics #6741 M) and Martin-Luther-University Halle-Wittenberg (Ethics #2019-103). Relevant clinical information of the patient cohort is included in Supplementary Data 1. Informed consent to participate in research was obtained from all patients or their legal guardians as stipulated by the study protocols.
TAM / ML-DS blast enrichment by flow cytometry
TAM and ML-DS samples obtained from the AML Berlin-Frankfurt-Münster (BFM) study group were FACS-sorted to obtain pure leukaemic blasts prior to the 10X workflow (Supplementary Fig. 10, Supplementary Data 1). Mononuclear cells were enriched with density gradient centrifugation and subsequently FACS sorted. Blast was defined as CD45dim/CD33 + /CD117 + /CD7 + /CD11a-.
10X single-cell RNA sequencing (scRNA-seq)
Fresh foetal livers were dissociated into single-cell suspensions. For the leukaemia samples (bone marrow aspirates or peripheral blood), peripheral blood mononuclear cells were prepared by density centrifugation using Lymphoprep (Stemcell) according to manufacturer’s instructions, also generating single-cell suspensions. The suspension was passed through a 70μm cell strainer (Falcon) and washed with PBS. If necessary, live cell enrichment using a Dead Cell Removal kit (Miltenyi Biotec) and red blood cell removal using the eBioscience 10X RBC Lysis Buffer (Multi-species) were performed as per the manufacturer’s instructions. For some foetal liver samples, cells are further sorted into CD45+ and CD45- fractions using magnetic beads (detailed in Supplementary Data 1). All enriched live cells were washed and counted using a hemocytometer with trypan blue; single cell suspensions were adjusted to 1000 cells/ul accordingly. Cells were loaded onto the Chromium 10X controller as per the standard protocol of the Chromium Single Cell 3’ Reagent kits (V3 chemistry) or 5’ Reagent kits (v2 chemistry) in order to capture between 7000 cells/chip position (detailed in Supplementary Data 1). All the following steps were performed according to the standard manufacturer protocol. Post GEM-RT clean-up, cDNA amplification, and 5′ gene expression library construction were carried out according to the manufacturer’s instructions. The resulting libraries were sequenced on the Illumina Novaseq 6000 platform, aiming for an average of 300,000 reads per cell.
Quality control and preprocessing of scRNA-seq data
Raw sequencing data were processed and mapped to the reference genome (GRCh38 1.2.0 or GRCh38 2020-A as detailed in Supplementary Data 1), using Cell Ranger pipeline62. The filtered count matrix, outputted by Cell Ranger, was then further QC’ed using Seurat (v4.0.1)63 in R (v4.0.4). Cells with <300 genes, <500 UMIs, or mitochondrial fraction exceeding 30% were removed. Scrublet (v0.2.3)64 was used to identify doublets. Cells were excluded if identified as doublets by Scrublet, or having a doublet score > 0.5. Ambient mRNA contamination was removed with SoupX (v1.6.1)65. High resolution clusters (resolution=10) with >50% cells failing QC were also excluded.
The data were log-normalised and scaled, and principal components were calculated using highly variable genes, following the standard Seurat workflow. Louvain clustering was performed (resolution = 1), and a uniform manifold approximation and projection (UMAP) was calculated, using the top 75 principal components. No integration or batch-correction methods were performed to preserve biological variation across different karyotypes and cancer samples.
Bulk mRNA sequencing and processing
RNA was isolated from cells using the Quick-RNA Miniprep Kit (Zymo Research). Paired-end libraries with 2 × 50, 75, 100 and 150 bp reads were prepared from the extracted RNA using the TruSeq Stranded total RNA LT Sample Prep (RiboZero Gold, Illumina) using Illumina Methodology by Novogene Company, Ltd. Raw sequencing reads were trimmed with fastp (v0.24.0)66 and aligned to GRCh38 using STAR (v2.6.0)67. Mapped reads were annotated using Ensembl v.108. Gene expression levels in transcripts per million (TPM) were quantified from the mapped reads using salmon (v1.10.3)68.
Whole genome sequencing
Whole genome sequencing (WGS) was performed on the foetal tissues with abnormal karyotypes, and those from two cases of aggressive ML-DS: one relapsed twice, and one refracted to standard chemotherapy (details in Supplementary Data 1). Bulk DNA sequencing was performed as previously described69. In brief, DNA was extracted using the AllPrep DNA/RNA/Protein Mini Kit (QIAGEN) following the standard protocol, and short insert (~500 bp) genomic libraries were generated. Finally, 150 bp paired-end sequencing was conducted on the Illumina NovaSeq 6000 platform according to Illumina’s standard library generation protocols (with PCR). The average sequence coverage was at least 30X per sample (details in Supplementary Data 1).
Raw DNA sequencing data were aligned to the GRCh38 (Ensembl 103) reference genome using the Burrows-Wheeler algorithm (BWA-MEM)70.
Sample-level genotyping
To ensure that sequencing data (both WGS and scRNA-seq) from the same individuals are correctly labelled, sample-level genotyping was performed using the matchBAMs function from the R package alleleIntegrator46.
The karyotype of each donor was also confirmed. For those with WGS data (see Supplementary Data 1), we compared the coverage of affected chromosome(s) to the rest of the genome as follows:
-
Single-chromosome trisomy: coverage of the trisomic chromosome is approximately 1.5 times higher than that of other disomic chromosomes (other copy number regions were excluded, if any).
-
Monosomy X: There is no coverage on chromosome Y (confirming that the individual is a female), and the coverage of chromosome X is half that of other disomic chromosomes.
-
Whole genome trisomy: We assessed the distribution of the alternate-allele frequency at heterozygous single-nucleotide polymorphisms (SNPs), which showed a bimodal distribution with peaks around 1/3 and 2/3.
For cases with only scRNA-seq data, we first combined all scRNA-seq samples from the same individual and identified the individual-specific heterozygous SNPs using the R package alleleIntegrator46. We then calculated the alternate/B-allele frequency (BAF) at heterozygous SNPs, aggregated across all high-quality sequencing reads (mapping quality >= 200, base quality >= 20, minVarQual = 225) on each chromosome. By assessing BAF distributions for each individual, we confirmed that trisomic chromosomes exhibit a bimodal BAF distribution with peaks around 1/3 and 2/3, while disomic chromosomes show a unimodal distribution around 0.5.
Cell type annotation
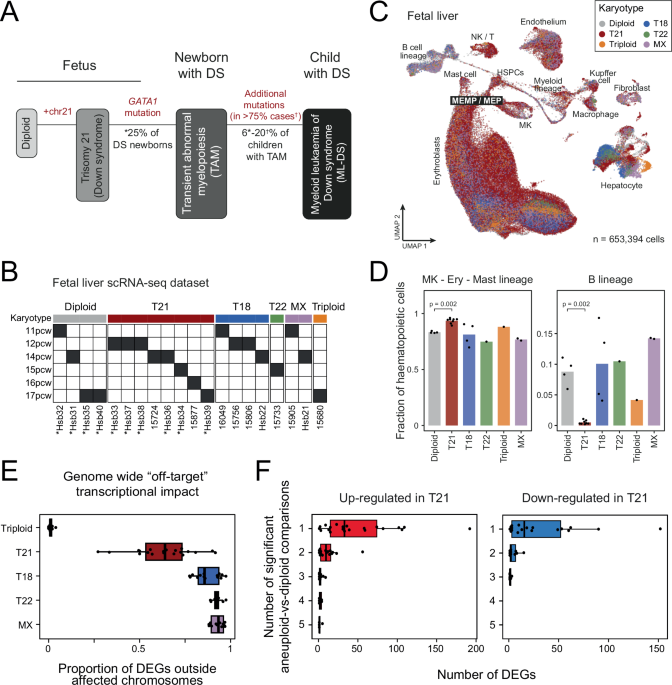
Foetal liver scRNA-seq dataset
A semi-automated cell type annotation using a label transfer approach was employed. Using Celltypist71, a logistic regression model was trained on the reference scRNA-seq atlas of the human foetal livers (Popescu, DM., Botting, R.A., Stephenson, E. et al., 201910). The model was then used to calculate predicted similarity scores for individual cells in our query foetal liver dataset against each reference class (i.e., different cell types in the reference dataset) (Supplementary Fig. 1D). Cells were assigned the label of the class with the highest positive similarity score. Annotation was further refined by manually assessing the expression of well-established cell-type-specific marker genes (Supplementary Fig. 1C) in high-resolution Louvain clusters.
To quantify the proportion of different haematopoietic lineages in the foetal livers of different karyotypes, we first excluded all non-haematopoietic cells, including: endothelial cells, fibroblasts, hepatocytes, cholangiocytes, mesothelial cells, neurons, and trophoblasts. The remaining cells were grouped by donor ID and corresponding lineages as follows:
-
HSC/MPP
-
Megakaryocyte – Erythroid – Mast cell lineage: MEMP/MEP, early megakaryocytes, megakaryocytes, early/mid/late erythroblasts, mast cells.
-
B lineage: LMPP/ELP, pro B cells, pre B cells, B cells.
-
Myeloid lineage: CMP/GMP, DC1, DC2, pDC, pro-monocytes, monocytes, macrophages, Kupffer cells, pro-myelocytes, myelocytes.
-
NK/T lineage: ILC precursors, natural killer/T cells.
The proportion of each lineage in each donor was calculated as the number of cells in the lineage divided by the total number of haematopoietic cells from that donor.
Comparisons of the lineage proportions between those with trisomy 21 samples and diploid foetal livers were performed using donor-level proportions with a one-tailed Wilcoxon rank-sum test (using the wilcox.test function in R), treating each donor as a biological replicate.
Leukaemia scRNA-seq dataset
A similar approach was used to annotate both leukaemia scRNA-seq datasets (TAM/ML-DS and other leukaemias), where automated label transfer was followed by manual assessment of canonical cell type-specific marker genes. However, as these datasets contain neoplastic cells that may not closely resemble normal cellular transcriptomes, we employed a logistic regression method previously detailed in Young et al., 201869. Specifically, a logistic regression model with elastic net regularisation (alpha = 0.99) was trained on our diploid foetal liver scRNA-seq data (reference dataset), using the cv.glmnet function from the glmnet R package. This model was applied to calculate the predicted similarity scores for individual cells in the leukaemia datasets against each reference class (i.e., different cell types in the reference dataset) (Supplementary Fig. 3C). To allow for the possibility that some query cells may not match strongly to any of the reference cell types, softmax normalisation was not used. Cells were assigned the label of the class with the highest positive similarity score. Annotation was further refined by manually assessing the expression of well-established cell type specific marker genes (Supplementary Fig. 3D) in high-resolution Louvain clusters.
Identification of GATA1 mutated neoplastic cells in TAM / ML-DS scRNA-seq data
Genotyping for the presence of GATA1 mutations in single-cell transcriptomes
The exact GATA1 mutation(s) acquired in each TAM / ML-DS case was obtained from clinical history and confirmed with our WGS analyses where possible. Next, we examined the scRNA-seq reads from each sample, using samtools72 to identify high quality reads overlapping the GATA1 mutation position +/− 70 bases. These reads were sorted into GATA1 mutant or wild-type reads based on the nucleotide sequence; and assigned to their corresponding cell barcodes. The number of mutant or wild-type GATA1 reads per cell was tallied. It is expected that only one allele of GATA1 is expressed in a given cell, as GATA1 is located on chromosome X outside the PAR region, combined with X-inactivation in females. Therefore, cells were classified as mutant (and therefore neoplastic cells) if they expressed only the GATA1 mutant allele, and normal if they exclusively expressed the wild-type allele. For cells with both wild-type and mutant reads (likely due to doublets or ambient RNA contamination), cells were classified as mutant if they had at least five more mutant reads than wild-type reads, or wild-type if the reverse was true. Otherwise, they were considered ambiguous.
Differential gene expression analysis
All differential gene expression analyses outlined below employed a donor-level pseudo-bulk approach. Donor-level pseudo-bulks were generated by aggregating raw counts per gene across all relevant cells from each donor. Pseudo-bulks with fewer than 30 cells were excluded, along with genes on chromosome Y. DESeq273 was used to perform differential gene expression analysis, following the workflow outlined in the vignette [https://bioconductor.org/packages/devel/bioc/vignettes/DESeq2/inst/doc/DESeq2.html]. In brief, a negative binomial generalised linear model was fitted to estimate the effect of independent variables on gene expression levels. Wald statistics with Benjamani-Hochberg correction for multiple hypothesis testing were used to identify genes with significant differential expression (i.e., genes with log2 fold-change (log2FC) of expression level significantly different from zero). The log2FC for each gene was extracted and subjected to shrinkage correction to improve accuracy for genes with high dispersion. Genes were identified as significantly differentially expressed (DEGs) using the following criteria unless stated otherwise:
-
1.
Adjusted p-value < 0.05
-
2.
Absolute log2FC >= 0.5
-
3.
Expressed in >= 10% of cells across all samples from either group
Transcriptional consequences of different abnormal karyotypes on foetal liver cell types
To assess the transcriptional impact of different abnormal karyotypes on foetal liver cell types, we performed pseudo-bulk differential expression analyses comparing each cell type from individual abnormal karyotypes with the diploid counterpart (Fig. 1E, F; Supplementary Fig. 2; Supplementary Data 2). Due to low cell numbers, some cell types were grouped together:
-
Megakaryocyte: early megakaryocytes and megakaryocytes.
-
Myelocyte: pro-myelocytes and myelocytes.
-
Monocyte: pro-monocytes and monocytes.
-
B cell progenitors (“B.cell.prog”): LMPP/ELP; pro B cells; and pre B cells.
-
NK/T: ILC precursors and natural killer / T cells.
To estimate the sample-specific size factor, the estimateSizeFactors function was used with controlGenes parameter specified to only genes on disomic chromosomes. For the triploid vs diploid comparison, controlGenes is specified as genes on chromosome X outside the PAR region (which is known to escape X-inactivation), with the assumption that despite being triplicated, the expression level of these genes is least likely to be affected compared to diploidy due to the X-inactivation mechanism in females. The model design used was “Gene_expression ~ Karyotype + Assay”.
Genes were considered to be expressed by a given cell type if they were expressed in >= 10% diploid cells of that cell type.
For each karyotype, the genome-wide “off-target” transcriptional impact in each cell type was defined as the proportion of DEGs residing outside the affected chromosome(s) (Fig. 1E).
Derivation of the trisomy 21 – leukaemia transcriptional module
We defined the “trisomy 21 – leukaemia” transcriptional module as the transcriptional changes observed in trisomy 21 foetal liver MEMP/MEP cells compared to their diploid counterpart that is contained within the ML-DS transcriptome.
First, the ML-DS transcriptome is defined as DEGs between diagnostic ML-DS blasts (from bone marrow samples only) compared to diploid foetal liver MEMP/MEP cells. To minimise potential confounding artifacts, only diploid foetal liver samples sequenced using the same 10X reagent kit (Chromium Single Cell 5’ Reagent kits – v2 chemistry) as the ML-DS samples were included. The model design used was “Gene_expression ~ Group”. We then examined which of these DEGs were already significantly perturbed in the same direction in trisomy 21 foetal liver MEMP/MEP. We performed a second pseudobulk analysis comparing expression levels of these genes in cells with trisomy 21 and diploid cells. Here, all relevant foetal liver samples sequenced with either 5’ or 3’ reagent kit were included. To account for the effects of different reagent kits, the model design used was “Gene_expression ~ Karyotype + Assay”, where “Assay” is a two-level factor representing reagent kits. Genes which met criteria (1) above were considered to be significantly differentially expressed in trisomy 21 foetal liver MEMP/MEP, and thus included in the “trisomy 21 – leukaemia” transcriptional module. Criteria (2) and (3) were excluded here as we wanted to retain all relevant changes in expression level, including very subtle perturbations represented by low log2FC values. Full results from both analyses can be found in Supplementary Data 3.
Derivation of the GATA1 – leukaemia transcriptional module
The “GATA1 – leukaemia” transcriptional module reflects the transcriptional consequences of GATA1 mutations observed in TAM blasts compared to trisomy 21 MEP cells, which are contained within the ML-DS transcriptome.
We first performed a pseudobulk differential expression analysis comparing diagnostic bone marrow ML-DS blasts with normal trisomy 21 MEP cells. More specifically, the latter cells were obtained from post-treatment remission bone marrow samples of four different ML-DS patients (L019, L038, L040, L041) with the highest number of captured MEP cells. DEGs were identified using all three criteria outlined above. We then tested for the differential expression of these genes between conventional (i.e. non-recurrent) TAM blasts and the same set of trisomy 21 MEP cells. Genes which met criteria (1) were considered to be significantly differentially expressed in TAM blasts as transcriptional changes induced by GATA1 mutations, and thus included in the “GATA1 – leukaemia” transcriptional module. Full results from both differential expression analyses can be found in Supplementary Data 4.
To better understand the “GATA1 -leukaemia” transcriptional module, we examined the expression of each gene across normal haematopoietic cells from our diploid foetal liver scRNA-seq dataset. The average expression per gene was calculated for each cell-type category using the AverageExpression function from the Seurat package63. Genes with no detectable expression in any haematopoietic cell categories were excluded from the heatmap shown in Supplementary Fig. 5A.
Derivation of the ML-DS leukaemia transcriptional module
The “ML-DS leukaemia” transcriptional module captures the transcriptional differences between the leukaemic state of ML-DS and the pre-leukaemic state of TAM blasts. A pseudobulk differential expression analysis was performed comparing ML-DS cancer cells from diagnostic bone marrow samples against conventional TAM blasts. Genes were considered to be DEGs if all three criteria outlined above were met. Full results can be found in Supplementary Data 5.
Grouping all ML-DS cases into one category may have obscured between-patient heterogeneity, which is known to be a prominent feature across cancer types. To address this, we performed a second analysis comparing the conventional TAM group against individual ML-DS cases. DEGs for each comparison were extracted using the above 3 criteria. This analysis revealed a large number of transcriptional perturbations in ML-DS compared to TAM which are individual-specific (Supplementary Fig. 5B).
Characterisation of the transcriptional signature of progressive ML-DS
To investigate the transcriptional changes potentially associated with progressive ML-DS and chemotherapy resistance, we specifically looked for markers associated with chemotherapy resistance in the progressive blasts in L076 and L038 independently, using the FindMarkers function from the Seurat package63. For L076, we compared progressive cancer cells from relapse 1 and relapse 2 against those from the initial diagnostic samples. For L038, we compared cancer cells with TP53 loss in timepoint 1 samples against cancer cells without TP53 loss in the initial diagnostic samples. Full results can be found in Supplementary Data 8.
Transcriptional module enrichment analysis
The trisomy 21 – leukaemic gene module was composed of all genes derived from the above analysis. The GATA1 – leukaemic gene module and ML-DS leukaemic gene module from the above analyses were further refined to retain only genes expressed in >= 20% more cells in the group with higher expression. The enrichment score for each gene module (composed of both up- and down-regulated genes) in single-cell RNAseq data was calculated using the function AddModuleScore_UCell from the R package UCell74 (v1.3.1). For the bulk transcriptome dataset, the enrichment score was computed using the function simpleScore from the R package singscore75 (v1.10.0).
Phylogenetic reconstruction of progressive ML-DS
Somatic variants calling
We investigated the genetic evolution of relapse and refractory ML-DS in child L076 and child L038 respectively. For both cases, the genetic phylogenetic relation was reconstructed based on somatic single-nucleotide variants (SNVs, also referred to as substitutions). SNVs in both the diagnostic and TP1 WGS samples were called using the CaVEMan algorithm76 (v1.18.2), and short insertions/deletions were called using Pindel algorithm77 (v3.10.0) in an unmatched analysis of each sample against an in silico normal human reference genome. In addition to the inbuilt QC filters, we further applied a series of stringent filters to remove low quality variants and germline variants, following the detailed workflow outlined in Coorens et al., 201945. Briefly, we only retained variants with a high median alignment score of supporting reads (ASMD > = 140), and required that fewer than half of the reads were clipped (CLPM = 0), as well as not falling within ten base pair of a deletion or insertion called by Pindel. We then recounted across both samples the variant allele frequency of all substitutions with a cut-off for base quality of 25 and read mapping quality of 30. Variants were also filtered out if they were called in a region of consistently low depth across both samples (excluding copy number segments). Next, to filter out germline substitutions, we fitted a binomial distribution to the combined read counts across all samples per SNV site, and applied a one-sided exact binomial test to calculate the probability that the SNV is consistent with being a germline variant. For the remaining variants, we calculated the site-specific error rates by interrogating the same sites in a panel of normal blood samples, using a beta-binomial model derived from the Shearwater variant call78. Variants which were indistinguishable from the background error rates were removed. All variants which passed all filters were visually inspected using the genome browser Jbrowse79 to exclude further sequencing or mapping artefacts. The final list of somatic substitutions can be found in Supplementary Data 6 for L076 and Supplementary Data 7 for L038.
Single-base substitution signature analysis
The single-base substitution profile of each WGS sample from L076 and L038 was further deconvolved into linear combinations of known COSMIC reference signatures (v3.4) using the R package mSigAct80 (v3.0.1). The set of reference signatures provided was: SBS1, SBS5, SBS13, SBS15, SBS18, SBS31, SBS35, SBS44, SBS84, and SBS86. Signature exposures per sample can be found in Fig. 4A, C.
WGS-derived copy number aberrations
We identified copy number aberrations in each WGS sample from L076 and L038 using the COBALT, AMBER, and PURPLE toolkit developed by the Hartwig Medical Foundation, following their documentation available on github:https://github.com/hartwigmedical/hmftools. Briefly, the BAF of heterozygous single-nucleotide polymorphism (SNP) sites was computed using AMBER (v3.9), and read depth ratios were calculated with COBALT (v1.14.1), both in “tumour-only” mode. BAF information and read-depth ratios were integrated to estimate the purity, ploidy and copy number profile of each tumour using PURPLE81 (v3.8.4).
Detection of copy number aberrations in scRNA-seq data
Using the R package alleleIntegrator45 (v0.9.1), we examined the copy number status of individual leukaemic blasts from patient L076 (relapse ML-DS), L038 (refractory ML-DS) and L067 (diploid MDS with GATA1 mutation). Heterozygous SNPs were identified from WGS data from each individual, and phasing of heterozygous SNPs across copy number segments with allelic imbalance was performed. The number of scRNA-seq reads supporting the major/minor allele across each copy number segment in each cell was calculated. Finally, the posterior probability of each cell harbouring the altered or normal copy number state for each copy number segment was calculated.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.