PIAS4 exhibits high expression levels in breast cancer and is inversely associated with prognosis
Comprehensive gene expression profiles of breast cancer and corresponding normal breast tissues were obtained from the TCGA database. We analysed the differential expression of these genes in breast cancer and normal breast tissues (Fig. 1A). Notably, we observed elevated expression levels of PIAS3 and PIAS4 in tumour tissues compared with their respective normal tissues (Fig. 1B). Kaplan–Meier overall survival analysis revealed that patients with higher PIAS4 expression exhibited shorter overall survival than their counterparts (Fig. 1C). In contrast, high PIAS3 expression showed no significant effect on overall survival. (Fig. 1D). Breast cancer can be classified into different molecular subtypes, commonly characterised by the expression of ESR1, PGR, and ERBB2. We explored the correlation between PIAS4 expression and these three markers and RB1 using TCGA database. The analysis indicated that PIAS4 expression showed only weak correlations with ESR1, PGR, ERBB2 and RB1 (Fig. S1A–D). We conducted a comparative analysis of PIAS4 protein levels using clinical tissue microarrays containing 45 normal tissue samples and 101 tumour samples. Our findings demonstrated a significantly elevated protein level of PIAS4 in tumour tissues compared to normal breast tissues (Fig. 1E, F). Finally, survival curves were generated using microarray-based PIAS4 protein data from 101 tumour tissues and their corresponding clinical data, which revealed a negative correlation between high PIAS4 protein expression and patient prognosis (Fig. 1G). We also examined the correlation between PIAS4 protein expression and ER, PR, and HER2 proteins in the above-mentioned tissue microarrays. The results revealed that PIAS4 protein levels were only weakly correlated with ER, PR, and HER2 expression (Fig. S1E–G). In summary, we observed high PIAS4 expression in breast tumour tissues that negatively correlated with prognosis.
Fig. 1: PIAS4 exhibits high expression levels in breast cancer and is inversely associated with prognosis.
A Differential gene expression was detected between normal and malignant breast tissues with |log2FoldChange | >2 and adjusted P value < 0.05, as obtained from the TCGA database. Notably, the expression patterns of PIAS4, PIAS3, and canonical breast cancer markers including BRCA1, BRCA2, ESR1, PGR, and ERBB2 were highlighted. B Expression profiles of the four members of the PIAS family examined using TCGA data. C, D Kaplan–Meier overall survival curves generated to predict clinical prognosis based on PIAS4 (C) and PIAS3 (D) expression levels. E Comparative analysis of PIAS4 expression between normal breast tissues and breast tumour tissues in clinical tissue microarrays. F Representative images of PIAS4 protein level in tumour and corresponding benign glandular tissues. G Correlation between prognosis and PIAS4 expression investigated in clinical tissue microarrays.
Inhibition of PIAS4 delays cell proliferation and cell cycle G1 phase progression
To further elucidate the role of PIAS4 in breast cancer, we used the functional annotation tool DAVID to categorise the 92 interacting proteins of PIAS4 previously identified [21]. Figure 2A illustrates the major biological functions based on the significance and number of proteins with enriched terms. Notably, transcription, regulation of transcription, and DNA binding had the highest significance in Gene Ontology term analysis (Fig. 2A, Fig. S2A). Gene Set Enrichment Analysis (GSEA) results indicated that silencing PIAS4 with specific siRNAs impeded cell cycle progression, accompanied by upregulation of a subset of cell cycle–related genes (Fig. 2B, C, Fig. S2B). Perturbation of PIAS4, either through knockout, specific siRNA/shRNA interference, or SUMOylation inhibitor treatment, markedly affected MEF and breast cancer cell MCF7 and BT-474 proliferation (Fig. 2D–F, S2C–S2E, and Fig. S3). Additionally, overexpression of Pias4 in Pias4-/- MEF cells restores cell growth (Fig. S2F).
Fig. 2: Inhibition of PIAS4 blocks cell cycle G1 phase progression and inhibits cell proliferation.
A Biological processes associated with PIAS4 were elucidated using the enrichment tool DAVID. B Enrichment analysis revealed PIAS4-targeted genes in the KEGG pathway via GSEA. C Enrichment analysis revealed PIAS4-knockdowned genes in the cell cycle via GSEA. D–F Interference of PIAS4 in PIAS4 + / + MEF (D), MCF7 (E), and BT-474 (F) cells was assessed using MTS assay to determine relative cell viability.
Therefore, we conducted cell cycle analysis on Pias4 + /+ and Pias4-/- MEF cells. Pias4 knockdown significantly delayed MEF in G1 phase (Fig. 3A, and Fig. S4A–S4B). Similarly, depletion of PIAS4 in breast cancer cells resulted in noticeable G1 phase delay (Fig. 3B, C, and Fig. S4C–S4D). To further elucidate the regulatory role of PIAS4 in cell-cycle progression, thymidine analogue EdU incorporation assays were performed to examine the effect of PIAS4 depletion on DNA replication. The results showed that PIAS4 siRNA treatment markedly reduced the number of MCF-7 and BT-474 cells entering the DNA replication phase (Fig. 3D, E).
Fig. 3: Inhibition of PIAS4 impairs cell cycle progression at the G1 phase.
A Flow cytometry was used to analyse and sort wild-type (PIAS4 + /+) and knockout (PIAS4-/-) MEF cells. B, C Depletion of PIAS4 in breast cancer cell lines MCF7 and BT-474 induced G1 phase delay, as determined by flow cytometry. D, E Cell proliferation of MCF7 and BT-474 cells was evaluated using a thymidine analogue incorporation assay. F, G MCF7 and BT-474 cells were transfected with PIP-Fucci plasmids to monitor cell cycle progression by flow cytometry following PIAS4 knockdown.
To monitor cell-cycle dynamics, a pLenti-PGK-Neo-PIP-FUCCI plasmid encoding mCherry-Geminin and mVenus-Cdt1 fusion proteins was used. In this system, green fluorescence (mVenus-Cdt1) indicates G1 phase, red fluorescence (mCherry-Geminin) represents S phase, and yellow fluorescence (co-expression of both) corresponds to G2/M phase. Flow cytometric analysis of FUCCI-reporter cells revealed that PIAS4 knockdown significantly increased the proportion of MCF-7 and BT-474 breast cancer cells in the G1 phase, accompanied by a marked reduction in S-phase cells (Fig. 3F, G). Thus, the deletion of PIAS4 disrupts cell cycle G1 phase progression and inhibits cell proliferation.
Inhibition of PIAS4 reduces the phosphorylation level of RB1 and the expression of its target genes
The cell cycle is tightly regulated by intricate mechanisms that ensure genomic stability and cell proliferation. During the G1/S checkpoint of the cell cycle, cyclin D-CDK4/6 and cyclin E-CDK2 phosphorylate the tumour suppressor protein RB1, leading to its inactivation. Hyperphosphorylation of RB1 leads to its dissociation from the E2F transcription factor, resulting in a relaxed chromatin structure and the transcription of genes regulated by RB1. Therefore, we investigated whether the inhibition of PIAS4 caused cell cycle G1 phase delay by modulating RB1 phosphorylation. Knockout or knockdown of PIAS4 in MEF and breast cancer cells resulted in the suppression of RB1 phosphorylation not only at the S807/811 sites but also at the S780 and S795 sites (Fig. 4A–D and Fig. S5A–D). Treatment with the SUMOylation inhibitor TAK-981 reduced RB1 phosphorylation in a dose-dependent manner (Fig. S5E, F). Conversely, overexpression of PIAS4 effectively restored RB1 phosphorylation in PIAS4 knockout MEF (Fig. 4E and Fig. S5G). Additionally, downregulation of total RB1 levels was observed in PIAS4-silenced cells (Fig. 4A–D). To evaluate whether PIAS4 regulates RB1 at the transcriptional level, we examined the correlation between PIAS4 and RB1 expression in TCGA datasets (Fig. S1D). Although the correlation was statistically significant (R = 0.305, P < 0.001), the relatively low correlation coefficient indicates a weak association between PIAS4 and RB1 mRNA expression. Therefore, PIAS4-mediated regulation of RB1 is likely to occur predominantly at the post-transcriptional or protein level rather than at the level of transcription.
Fig. 4: Inhibition of PIAS4 decreases the phosphorylation level of RB1 and the expression of its target genes.
A, B Western blot was used to detect RB1 phosphorylation in Pias4 + /+ MEF cells with Pias4 knockout and knockdown. C, D Western blot was performed to measure the cell cycle-related proteins after treatment with specific PIAS4 siRNA in breast cancer MCF7 and BT-474 cells. E RB1 phosphorylation in PIAS4-/- MEF cells overexpressing PIAS4 measured using western blot. F, G Western blot analysis was performed to examine CDK6 protein stability in MCF7 and BT-474 breast cancer cells following PIAS4 knockdown and cycloheximide treatment. H–K qRT-PCR was used to determine the expression levels of RB1 target genes after PIAS4 knockdown or inhibition in breast cancer MCF7 and BT-474 cells. Data are presented as the means ± SD from three independent experiments.
In both PIAS4 knockout and knockdown cells, we observed a moderate increase in CDK6 expression, whereas cells with high PIAS4 expression exhibited a marked reduction in CDK6 levels (Fig. 4A–E). To further investigate whether PIAS4 affects the stability of CDK6 protein, we examined CDK6 protein turnover following PIAS4 depletion. The results showed that PIAS4 knockdown significantly enhanced the stability of the CDK6 protein (Fig. 4F–G and Fig. S5H–S5I). In addition to regulating CDK6 protein stability, we asked whether PIAS4 also affects CDK6 at the transcriptional level. We first analysed the association between PIAS4 expression and CDK6 mRNA levels using TCGA breast cancer datasets (Fig. S5J) and observed a weak but statistically significant positive correlation (P < 0.001, R = 0.214). Consistent with this observation, PIAS4 knockdown resulted in a modest increase in CDK6 mRNA expression in breast cancer cell lines MCF7 and BT-474 (Fig. S5K, L). These data indicate that PIAS4 depletion leads to elevated CDK6 mRNA levels and enhanced CDK6 protein stability, together contributing to the increased total CDK6 protein abundance observed upon PIAS4 knockdown.
Depletion of PIAS4 using a specific siRNA or SUMOylation inhibitor TAK-981 led to reduced expression of RB1 target genes, such as CDK1, E2F1, CCNE2, and CCNA2 (Fig. 4H–K). These findings suggest that PIAS4 regulates both RB1 phosphorylation and expression of its target genes.
To determine whether PIAS4-mediated regulation of RB1 phosphorylation and the transcription of RB1 downstream genes depends on cell-cycle progression, we first synchronised cells at the G1 phase by serum starvation. However, serum starvation alone produced only a modest effect, resulting in a slight increase in the proportion of G1-phase cells (Fig. S6A–D). In contrast, PIAS4 knockdown further increased the G1-phase population (Fig. S6A–S6D), reduced RB1 phosphorylation levels (Fig. S6E, F), and decreased the transcription of RB1 target genes (Fig. S6H, J).
We next synchronised cells at the G2/M phase using nocodazole treatment, which effectively increased the G2/M population by 30-50% (Fig. S6A, D). Under these conditions, PIAS4 knockdown again led to an increased proportion of G1-phase cells (Fig. S6A–D), reduced RB1 phosphorylation (Fig. S6E, F), and decreased expression of RB1 downstream genes (Fig. S6G, I).
These experiments indicate that PIAS4 regulates RB1 phosphorylation, the expression of RB1 target genes, and G1-phase progression even under conditions of serum starvation or nocodazole-induced synchronisation.
CDK6 mediates PIAS4 regulation of RB1 phosphorylation and cell cycle G1 phase progression
Considering the essential roles of CDK4/CDK6 and CDK2 in the cell cycle at the G1/S checkpoint, we investigated the interactions between PIAS4, CDK4/CDK6, and CDK2. Our findings indicate that PIAS4 specifically interacts with CDK6 and CDK4 but not with CDK2 (Fig. 5A). Consistent with this, Flag-CDK6 was found to interact with HA-PIAS4 (Fig. 5B). We performed a proximity ligation assay (PLA) to examine the endogenous interaction between CDK6 and PIAS4. Distinct PLA signals were detected, indicating close spatial proximity between CDK6 and PIAS4 (Fig. 5C, D). Notably, the majority of these fluorescent signals were localised within the nucleus, supporting the presence of a nuclear proximal interaction between CDK6 and PIAS4.
Fig. 5: PIAS4 interacts with CDK6 and regulates CDK6-mediated RB1 phosphorylation and cell cycle progression.
A, B Co-IP assays showing the interaction between HA-PIAS4 and either endogenous CDK6 or exogenous Flag-CDK6. C, D A proximity ligation assay (PLA) was performed to investigate the interaction between PIAS4 and CDK6 in breast cancer cell lines MCF7 and BT-474. E, F Western blot analysis showing the effect of CDK6 overexpression on PIAS4-mediated RB1 phosphorylation in breast cancer cells MCF7 and BT-474. G, H Flow cytometry analysis demonstrating the impact of CDK6 overexpression on PIAS4-mediated cell cycle G1 phase progression in breast cancer cells MCF7 and BT-474.
CDK6 overexpression partially rescued the downregulation of RB1 phosphorylation induced by PIAS4 knockdown in breast cancer cells (Fig. 5E, F). PIAS4 knockdown led to cell cycle G1 phase delay, whereas CDK6 overexpression partially rescued the cell cycle G1 phase delay induced by PIAS4 inhibition (Fig. 5G, H). These experiments confirmed that CDK6 interacts with PIAS4 within the nucleus and mediates PIAS4 regulation of RB1 phosphorylation and cell cycle G1 phase progression.
PIAS4-mediated SUMOylation modification of CDK6 promotes RB1 phosphorylation and cell cycle progression
PIAS4 regulates RB1 phosphorylation and cell cycle progression via CDK6. It possesses SUMO E3 ligase activity, conjugating SUMO1, SUMO2, and SUMO3 to substrate proteins, thereby regulating various biological processes. Therefore, we investigated whether PIAS4 SUMOylates CDK6 to regulate RB1 phosphorylation and G1 phase progression. Our findings revealed that CDK6 undergoes SUMOylation by SUMO1, SUMO2, and SUMO3 (Fig. 6A). Conjugation of SUMO molecules to target proteins depends on two C-terminal glycine residues. Thus, mutating these two glycine residues prevents the SUMOs from conjugating to the substrate. Mutating the two glycine residues caused the SUMO modification bands to disappear, confirming the specificity of these SUMOylation modifications (Fig. 6B).
Fig. 6: PIAS4-mediated SUMO3 modification of CDK6 promotes RB1 phosphorylation and cell cycle progression.
A Western blot was performed to assess whether CDK6 is modified by SUMO1, SUMO2, and SUMO3. B Western blot revealed changes in SUMOylation modification bands after mutating the double glycine residues of SUMO molecules. C Western blot analysis of the impact of wild-type, enzyme activity mutants (PIAS4-RINGΔ and PIAS4-W363A), and SUMO-interacting motif (SIM) mutant (PIAS4-SIM/AA) on CDK6 SUMOylation modification. D Four software tools predicted potential SUMO3 modification sites on CDK6. E Western blot analysis was conducted to evaluate the effects of mutations at predicted lysine sites on CDK6 SUMO3 modification. F Western blot was conducted to examine the impact of CDK6-6KR and CDK6-7KR mutations on CDK6 SUMO3 modification. G, H Western blot assessed the effect of CDK6-6KR and CDK6-7KR mutations on RB1 phosphorylation in breast cancer MCF7 and BT-474 cells. I, J Flow cytometry was used to analyse the impact of CDK6-6KR and CDK6-7KR mutations on the cell cycle in breast cancer MCF7 and BT-474 cells.
We further examined whether PIAS4 was essential for CDK6 SUMO modification. Overexpression of wild-type PIAS4 significantly increased CDK6 SUMO modification (Fig. 6C). Conversely, CDK6 SUMOylation modification decreased significantly when PIAS4 E3 ligase activity mutants, PIAS4-RINGΔ or PIAS4-W363A, were transfected (Fig. 6C), indicating that the E3 ligase activity of PIAS4 is crucial for CDK6 SUMOylation. Most SUMO E3 ligases possess a SUMO-interacting motif (SIM) that mediates non-covalent interactions with SUMO molecules. The SIM domain plays a crucial role in positioning the donor SUMO from the E2-SUMO complex, facilitating efficient SUMO transfer and catalysis [23]. Transfection with the PIAS4 SIM mutant, PIAS4-SIM/AA, showed only a subtle effect on CDK6 SUMOylation, suggesting that the SIM domain of PIAS4 is not critical for CDK6 SUMOylation (Fig. 6C).
Using four SUMO prediction tools, we identified 12 potential SUMOylation sites in CDK6 (Fig. 6D). Additionally, in vivo experiments confirmed that SUMOylation sites K29, K147, K160, K216, K230, K274, and K264 of CDK6 were highly SUMOylated (Fig. 6E). Mutating the SUMOylation sites K29, K147, K160, K216, K230, and K274 of CDK6 to arginine produced CDK6-6KR, and further mutation of K264 to arginine yielded CDK6-7KR. Compared with wild-type CDK6, the expression of mutants CDK6-6KR and CDK6-7KR in HEK-293T cells showed a significant reduction in the CDK6 SUMOylation modification band (Fig. 6F). Furthermore, transfection with wild-type CDK6 significantly promoted RB1 phosphorylation. Conversely, CDK6-6KR and CDK6-7KR transfection did not significantly enhance RB1 phosphorylation (Fig. 6G, H). Cell cycle analysis revealed that compared to wild-type CDK6, transfection with CDK6-6KR and CDK6-7KR no longer promoted cell cycle G1 phase progression. (Fig. 6I, J).These findings demonstrate that CDK6 undergoes both SUMO1 and SUMO2/3 modifications, which enhance CDK6-mediated RB1 phosphorylation, thereby promoting cell cycle G1 phase progression.
As mentioned above, PIAS4 is exclusively localised in the nucleus, whereas CDK6 is primarily cytoplasmic, with only a small fraction present in the nucleus. We therefore propose that PIAS4 interacts with and regulates the nuclear fraction of CDK6 (Fig. 5C, D). During the SUMOylation studies, both CDK6 and PIAS4 were overexpressed, which raised the concern that protein overexpression might alter their subcellular localisation and lead to a false-positive interaction.
To address this possibility, we performed validation experiments using mCherry-tagged PIAS4 and GFP-tagged CDK6 in breast cancer cells. The fluorescence results showed that overexpressed mCherry-PIAS4 remained strictly nuclear, while GFP-CDK6 displayed a distribution pattern consistent with that of endogenous CDK6 (Fig. 7C, D and Fig. S7). These findings confirm that the observed PIAS4–CDK6 interaction is genuine and not an artefact caused by protein overexpression.
Fig. 7: Inhibition of PIAS4 does not alter CDK6 subcellular localisation but decreases CDK6 kinase activity.
A, B Immunofluorescence staining was used to visualise the localisation of CDK6 (green), nucleus (blue), and the merged images (cyan) after treatment with PIAS4 siRNA in breast cancer MCF7 and BT-474 cells. C, D Western blot was performed to assess the protein level of CDK6 in the cytoplasm and nucleus after treatment with PIAS4 siRNA in breast cancer MCF7 and BT-474 cells. E Alignment of CDK6 protein sequences surrounding CDK6 Y24 residues of Homo sapiens, Mus musculus, Bos taurus, Gallus gallus, Danio rerio, Xenopus laevis, and Bactrocera dorsalis. F, G Western blot was performed to evaluate the phosphorylation of CDK6 Y24 after treatment with the PIAS4 siRNA in breast cancer MCF7 and BT-474 cells. H, I CDK6 IP kinase activity assays were performed to evaluate CDK6 activity following PIAS4 knockdown in MCF7 and BT-474 cells. J, K CDK6 IP kinase activity assays were conducted to assess CDK6 activity under PIAS4 knockout and overexpression conditions in PIAS4⁺/⁺ and PIAS4⁻/⁻ cells.
Inhibition of PIAS4 does not alter CDK6 subcellular localisation but decreases CDK6 kinase activity
We evaluated how PIAS4-mediated SUMOylation of CDK6 regulates cell cycle G1 phase progression. As a SUMOylation E3 ligase, PIAS4 regulates protein stability, localisation, activity, and interactions [21]. Our experiments demonstrated that abolishing PIAS4 with siRNA increases CDK6 stability (Fig. 4F, G and S5H, I). This observation is consistent with the increased levels of CDK6 protein in breast cancer cells following treatment with CDK4/6 inhibitors [24, 25].
We investigated whether PIAS4 inhibition affected CDK6 subcellular localisation. Using indirect immunofluorescence staining and nuclear and cytoplasmic extraction experiments, we observed that the fluorescent signal of CDK6 was distributed throughout the cells (Fig. 7A, B and Fig. S8A, B). The subcellular localisation of CDK6 did not change in response to PIAS4 siRNA or the SUMOylation inhibitor TAK-981 (Fig. 7A, B and Fig. S8A, B). Additionally, the cytoplasmic and nuclear components were isolated from breast cancer cells. Consistent with the immunofluorescence staining results, nuclear-cytoplasmic fractionation experiments indicated that the subcellular localisation of CDK6 remained unchanged after PIAS4 knockdown or inhibition. (Fig. 7C, D and Fig. S8C, D). These results indicate that knocking down PIAS4 does not alter CDK6 protein subcellular localisation.
CDK6 kinase activity is regulated by phosphorylation; hence, we examined the changes in CDK6 phosphorylation levels following PIAS4 inhibition. In breast cancer cell lines, the expression levels of two phosphorylation sites, tyrosine 13 (Y13) and threonine 177 (T177), necessary for CDK6 activation, were below the detection limit (data not shown). This finding is consistent with that of a previous study [26]. Additionally, we examined the CDK6 inhibitory phosphorylation site Y24. Conservation analysis of the CDK6 Y24 site and its adjacent amino acids across different species revealed that CDK6 Y24 and its neighbouring regions are highly conserved in mammals, birds, and zebrafish but less conserved in more distant related species, such as the African clawed frog and fruit fly. (Fig. 7E). Further experiments revealed that PIAS4 inhibition or the SUMOylation inhibitor TAK-981 treatment increased total CDK6 phosphorylation at the inhibitory site, Y24 (Fig. 7F, G and Fig. S8E, F). Furthermore, we employed a phosphorylation-deficient CDK6 mutant (Y24F) to mimic the dephosphorylated state at the Y24 residue. We found that expression of CDK6-Y24F markedly enhanced RB1 phosphorylation (Fig. S9A, B). These results indicate that increased phosphorylation at the CDK6 Y24 site plays an important inhibitory role in CDK6 activity and may contribute to the reduced RB1 phosphorylation observed upon PIAS4 knockdown.
To obtain robust evidence for the regulation of CDK6 by PIAS4, a CDK6 IP kinase activity assay was performed. The results showed that, despite an increase in total CDK6 protein following PIAS4 depletion, CDK6 kinase activity was reduced—consistent with the corresponding Western blot results showing decreased RB1 phosphorylation (Fig. 7H–J). Conversely, in PIAS4⁻/⁻ MEFs reconstituted with PIAS4, CDK6 protein levels were modestly decreased, whereas CDK6 kinase activity increased, in agreement with the elevated RB1 phosphorylation observed in the Western blots (Fig. 7K). To investigate the mechanism by which PIAS4 regulates CDK6 activity, we focused on cyclin D family proteins, which are essential regulators of CDK6 kinase activity. We therefore examined cyclin D1 protein levels in breast cancer cell lines following PIAS4 knockdown (Fig. 4C, D; Fig. 7H–K). The results showed that cyclin D1 protein levels were reduced upon PIAS4 depletion, whereas PIAS4 overexpression led to increased cyclin D1 expression. Moreover, CDK6 immunoprecipitation kinase assays revealed that the amount of cyclin D1 associated with CDK6 was also decreased in PIAS4-knockdown cells (Fig. 7H–I). Given the critical role of cyclin D1 in regulating CDK6 activity, we performed rescue experiments under PIAS4 knockdown conditions by overexpressing CDK6 alone or in combination with cyclin D1. Compared with CDK6 overexpression alone, co-expression of CDK6 and cyclin D1 more effectively rescued the reduction in RB1 phosphorylation caused by PIAS4 depletion (Fig. S9C–S9D). Collectively, these findings indicate that PIAS4 promotes CDK6 kinase activity by maintaining cyclin D1 protein levels and facilitating the interaction between cyclin D1 and CDK6, thereby regulating downstream RB1 phosphorylation and cell cycle progression.
PIAS4 and CDK6 inhibition synergistically suppress breast cancer growth
CDK4/6, crucial drivers of the cell cycle G1/S phase checkpoint, play a pivotal role in the development and progression of malignant tumours. Despite the significant therapeutic effects of pharmacological CDK4/6 inhibitors on numerous tumours, many patients still experience tumour progression. Hence, investigating the combination of CDK4/6 inhibitors with other targeted drugs is important for breast cancer treatment. Our findings indicate that PIAS4 is highly expressed in breast cancer and negatively correlates with prognosis. PIAS4 promotes G1 phase progression by regulating CDK6 activity, raising the question of whether PIAS4 inhibition can be combined with CDK4/6 inhibition for breast cancer treatment.
In breast cancer cells, the SUMOylation inhibitor, TAK-981, enhanced the inhibitory effects of CDK4/6 inhibitors on breast cancer growth (Fig. 8A–C). In the mouse xenograft tumour model, the combination of PIAS4 and CDK6 inhibition had no effect on mice’s body weight (Fig. 8D). However, PIAS4 knockdown enhanced the inhibitory effect of CDK6 silencing in breast cancer xenografts (Fig. 8E, F). Therefore, these experiments confirmed that PIAS4 inhibition synergises with CDK6 inhibition to suppress breast cancer development, providing valuable insights into breast cancer treatment.
Fig. 8: PIAS4 and CDK6 inhibition synergistically suppress breast cancer growth.
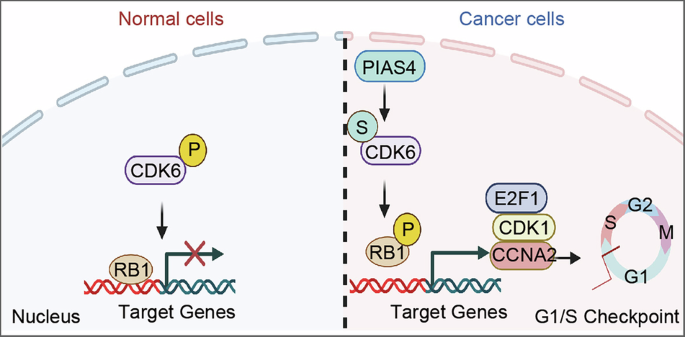
A–C Cell viability assays demonstrating the synergistic inhibitory effect of CDK4/6 inhibitors and SUMOylation inhibitors on breast cancer cell growth. D The impact of PIAS4 and/or CDK6 knockdown on mice body weight. E The impact of PIAS4 and/or CDK6 knockdown on mice tumour growth. F Representative luciferase bioluminescence images showing the effect of PIAS4 and/or CDK6 knockdown on breast mice tumour growth. G Schematic model illustrating the high expression of PIAS4 in tumour cells, which inhibits the phosphorylation of the CDK6 inhibitory site Y24 through SUMO modification. This promotes the phosphorylation of RB1 by CDK6, leading to the dissociation of transcription factor RB1 from the E2F1 complex, thereby enhancing the transcription of RB1 target genes and promoting cell cycle G1/S checkpoint progression.