Patients and specimens
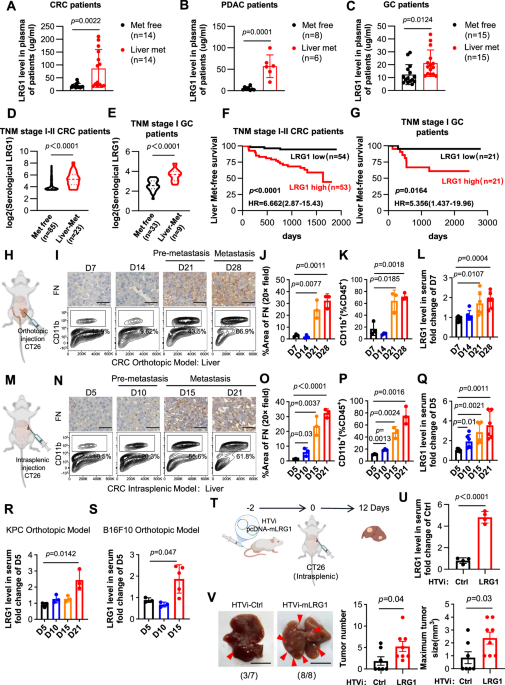
Serum samples from patients were obtained from BioBank, The Six Affiliated Hospital, Sun Yat-sen University. Paraffin-embedded human liver metastasis sections were obtained from the Sixth Affiliated Hospital of Sun Yat-sen University. The samples were used with informed consent under a protocol approved by the Medical Ethics Committee of the Sixth Affiliated Hospital of Sun Yat-sen University.
Animal experiments
Lrg1-flox mice (strain no. T009577) and Alb-Cre mice (strain no. T017784) on a C57BL6/J background were purchased from GemPharmatech (Nanjing, China). Lrg1wt/wt; Alb-cre (termed Lrg1(+/+)Hep), Lrg1wt/fl; Alb-cre (termed Lrg1(+/∆)Hep), and Lrg1fl/fl; Alb-cre (termed Lrg1(∆/∆)Hep) were generated by crossing Lrg1-flox mice and Alb-Cre mice. Genotyping of mice was performed by polymerase chain reaction (PCR) analysis of genomic DNA extracted from mouse tails using primers (Table S1).
For all tumor models, BALB/c or C57BL/6 J mice between 6 and 10 weeks of age were purchased from Guangdong GemPharmatech unless otherwise indicated. Mice of similar sex, age and weight were randomized before tumor inoculation. To determine the primary source of LRG1 in tumor models, mice were orthotopically implanted with 1×106 CT26 cells or MC38 cells, 5×104 KPC cells, or 1×106 B16F10 cells and intrasplenically implanted with 5×104 CT26 cells. The mice were euthanized at the indicated time points, and serum, specific organs and cells were collected to detect the expression of LRG1.
To detect liver metastases in Lrg1(+/+)Hep, Lrg1(+/∆)Hep and Lrg1(∆/∆)Hep mice, mice were implanted with 1×105 MC38 cells and euthanized. To determine the effect of PMN on liver metastases in Lrg1(+/+)Hep and Lrg1(∆/∆)Hep mice, mice were first orthotopically implanted with 1×106 MC38 cells. Afterward, the luciferase-labeled MC38-luc cells were intrasplenically implanted (1×105 cells for each mouse). The mice were euthanized, and the liver was rapidly harvested for ex vivo BLI.
To determine the effects of neutrophil depletion and DNase I on liver metastases in Lrg1(∆/∆)Hep mice, the mice were orthotopically implanted with 1×106 MC38 cells. On day 14, the mice received an initial i.p. injection of anti-Ly6G antibody (200 μg/mouse; Bio Xcell, #BE0075) every 3 days and an i.p. injection of DNase I (5 mg/kg, daily shots for 7 days from day 14, and then maintenance shots every 3 days). On day 40, the mice were euthanized, and the livers were rapidly harvested for ex vivo BLI. To test whether macrophages are the main contributors to IL6, mice were orthotopically implanted with 1×106 MC38 cells. On day 10, the mice received initial i.p. injections of clodronate liposomes (200 μl/mouse; Yeasen Biotechnology, 40337ES08) or liposomes (200 μl/mouse; Yeasen Biotechnology, 40338ES08) every 3 days and were euthanized on day 19 for further experiments.
To determine the therapeutic efficacy of anti-IL6 antibody in the CRCLM model, mice were i.p. injected with anti-IL6 antibody (200 μg/mouse; Selleck, A2118) or IgG antibody on day 10 after orthotopic injection, followed by every 3 days of injection. On day 14, MC38-luc cells were intrasplenically implanted; on day 32, the mice were euthanized, and the livers were rapidly harvested for ex vivo BLI. To determine the therapeutic efficacy of anti-LRG1 (C4, sc-390920) in a CRC intrasplenic model, mice were i.p. injected with anti-LRG1 (2 µg/mouse) 1 day before intrasplenic injection of cancer cells, followed by every 3 days of injection of the reagents for 21 days. On day 21, the mice were euthanized to detect liver metastases. To determine the therapeutic efficacy of AAV8-TGB-shLRG1 (5’-TGTCCATCTGTCGGTGGAATT-3’) in the CRCLM model, mice were i.v. injected with AAV-TGB-shLRG1 1 day before the orthotopic injection of MC38. On day 14, MC38-luc cells were intrasplenically implanted; on day 32, the mice were euthanized, and the livers were rapidly harvested for ex vivo BLI.
To determine the combined effect of anti-LRG1 and anti-PD-1 immunotherapy, BALB/c mice were used, and 2.5×105 CT26-luc cells in 25 μl of PBS were injected into the left main lobe of the mouse liver. On day 5, the mice were grouped according to ex vivo BLI and received an initial i.p. injection of anti-LRG1 every 2 days and anti-PD-1 (100 μg/mouse) every 3 days until completion of the experiment.
The mice were given a standard diet, allowed free access to water, and housed under a 12-h light/dark cycle. All the animal experiments were approved by the Institutional Animal Care and Use Committee of Sun Yat-sen University and conformed to the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health (National Academies Press, 2011) in China.
Ex vivo bioluminescence imaging (BLI)
For the liver metastasis models used in the present study, luciferase activity in the liver was used to monitor liver metastasis progression. The mice were i.p. injected with 100 μl of D-luciferin (150 mg/kg) and anesthetized for ex vivo BLI. The mice were euthanized, and the livers were rapidly dissected and placed in a plate filled with 2 ml of D-luciferin (150 mg/ml; diluted in PBS) for ex vivo BLI. BLI results were obtained using the Xenogen IVIS system. Light emission from the region of interest was quantified as photons/second/cm2/steradian (p/sec/cm2/sr) through Living Images software.
HTVi animal experiment
All these experimental procedures were approved by the Institutional Animal Care and Use Committee of Sun Yat-sen University. To overexpress LRG1 or IL6 in the liver, we hydrodynamically injected plasmid DNAs into the tail vein of mice following a previously published protocol. Each mouse received 10% of its body weight of saline containing 25–50 μg of plasmid DNA (pcDNA3.4-mLRG1 or pcDNA3.4-mIL6). The mice were maintained on a standard diet and sacrificed at the indicated time points.
Cell culture
CT26 colon adenocarcinoma cells and B16F10 melanoma cells were obtained from the American Type Culture Collection. MC38 colon adenocarcinoma cells were purchased from Kerafast. The murine pancreatic tumor KPC cell line was derived from the pancreatic tumors of KrasG12D/+; Trp53R172H/+; Pdx1-Cre C57BL/6 mice. The human CRC cell lines DLD1 and HCT116 were obtained from the American Type Culture Collection. The murine hepatocyte AML12 cell line and HL60 cell line were obtained from the American Type Culture Collection. CT26 cells were cultured in RPMI-1640 media supplemented with 10% fetal bovine serum and penicillin/streptomycin. MC38, KPC, DLD1 and HCT116 cells were cultured in DMEM supplemented with 10% fetal bovine serum and penicillin/streptomycin. AML12 cells were cultured in DMEM/F12 supplemented with 10% fetal bovine serum, 1× ITS (Sigma‒Aldrich), 40 ng/ml dexamethasone (Sigma‒Aldrich) and penicillin/streptomycin. HL60 cells were cultured in IMDM supplemented with 10% fetal bovine serum and penicillin/streptomycin. To differentiate HL60 cells into neutrophil-like cells, the cells were incubated in culture medium supplemented with 1% DMSO for 7 days. All the cells were cultured in a humidified incubator at 37 °C with 5% CO2.
RNA extraction and quantitative real-time PCR (qRT‒PCR)
Total RNA was extracted from cells and tissues using TRIzol reagent (Invitrogen) and ethanol precipitation. RNA reverse transcription was performed using a KAPA SYBR® FAST Universal kit. qPCR was subsequently conducted using a Roche Light-Cycler 480. The primers used are listed in Table S1.
Plasmid construction and lentiviral vector transduction
To generate the LRG1 overexpression plasmids, the full-length cDNA of mouse LRG1 was cloned and inserted into the pLenti-CMV vector. sgRNAs targeting CCDC25 were synthesized and cloned and inserted into Lenti-CRISPR-V2-Puro plasmids. shRNAs targeting TGFBR2 and TGFBR1 were synthesized and cloned and inserted into PLKO.1 plasmids. For stable transfection, the above constructs were cotransfected with pMD2. G, pRSV-REV and pMDLg/pRRE packaging plasmids were added to HEK293T cells in accordance with the manufacturer’s protocols. Lentiviral vector-containing supernatants were collected and used to knock out CCDC25 in DLD1 and HCT116 cells. The primers used are listed in Table S2.
Western blot
Cells and tissues were lysed in radioimmunoprecipitation assay (RIPA) buffer (Sigma‒Aldrich) supplemented with protease inhibitor (Roche) and quantified using a BCA kit (Thermo Scientific). The proteins were separated by SDS‒PAGE and transferred to NC membranes. The membranes were subsequently blocked with milk, followed by incubation with specific primary antibodies against LRG1, H3cit, p-AKT, t-AKT, p-ERK, t-ERK, p-p38, p38, p-SMAD2/3, t-SMAD2/3, p-SMAD1/5, t-SMAD1/5, p-PI3K, PI3K, TGFBR2, TGFBR1, CCDC25 and β-actin overnight at 4 °C. The details of the antibodies used are listed in Table S3. Afterward, the membranes were washed with TBST and incubated with HRP-conjugated secondary antibodies (1:5000; Sigma). The bands were visualized by chemiluminescence using the StarSignal Western ECL Substrate (GeneStar).
Immunofluorescence
The tissue was first fixed in 4% paraformaldehyde, embedded in paraffin and sectioned at a thickness of 4 μm. Then, the paraffin-embedded tissue sections were deparaffinized, rehydrated and subjected to antigen retrieval in EDTA buffer in a microwave. The sections were blocked with 5% BSA for 30 min at room temperature. The samples were subsequently incubated with specific primary antibodies against CD11b, LRG1, MPO, iNOS, and H3cit overnight at 4 °C. After rinsing with PBS, fluorochrome-conjugated secondary antibodies were added and incubated for 1 h at room temperature. The slides were counterstained with DAPI (D1306; Invitrogen). The details of the antibodies used are listed in Table S3. Observation and imaging were performed with a confocal microscope (Cell Observer; ZEISS, Germany). The analysis of fluorescence images relies primarily on ImageJ.
IHC staining
Paraffin-embedded tissue sections were deparaffinized with dimethylbenzene, dehydrated in an ethanol gradient, and subjected to antigen retrieval with EDTA buffer. Next, the tissues were blocked with normal goat serum and incubated with specific primary antibodies against fibronectin, LRG1 and p-Stat3 overnight at 4 °C. The tissue sections were then incubated with secondary antibodies for 1 h, and positive staining was visualized with an HRP DAB substrate kit, and the nuclei were counterstained with hematoxylin. The expression of LRG1 was quantified on the basis of the intensity of staining and the percentage of positive cells. In brief, the proportion of positive cells was estimated and given a score ranging from 1 to 4 (1, less than 5%; 2, 5–25%; 3, 26–50%; 4, > 51%). The average intensity of the positively stained cells was also given a score on a scale from 1 to 4 (1, no staining; 2, weak staining; 3, moderate staining; 4, strong staining). The final IHC score of each tissue sample was then calculated by multiplying the positive percentage score by the intensity score.
For LRG1, MPO, H3cit, IL6, F4/80, a-SMA, CD31, panCK, CD8a, and GZMb, tissue staining was performed with TSA (tyramide signal amplification) according to the manufacturer’s instructions. The slides were counterstained with DAPI (D1306; Invitrogen). The details of the antibodies used are listed in Table S3.
Tissue interstitial fluid collection
Tissue interstitial fluid from normal liver, PMN-liver and tumor tissue was isolated. Briefly, the tissues were chopped into pieces, and the tissue pieces were placed on a 70 µM filter on top of a 50 ml conical tube, followed by centrifugation at 200 × g for 10 min. The flowthrough tissue interstitial fluid was flash-frozen at -80 °C.
ELISA
ELISA kits were used to measure the levels of human LRG1 (Ray Bio), human IL6 (Boster, EK0410), mouse LRG1 (ELK Biotechnology), and mouse IL6 (Boster, EK0411) in cell culture supernatants or serum samples according to the manufacturer’s instructions. We detected plasma MPO-DNA using a previously described sandwich ELISA method. Briefly, 96-well microtiter plates were coated with 5 µg/ml anti-MPO monoclonal antibody (R&D, AF3667) as the capture antibody overnight at 4 °C. After blocking with 1% BSA buffer for 1 h, 50 µl of sample was added per well and incubated for 2 h at room temperature. A Quant-iTTM PicoGreenTM dsDNA Reagent (Thermo Fisher, P7589) and a kit were used to measure the levels of MPO-DNA following the manufacturer’s instructions. The absorbance was measured using a microplate reader.
Tissue dissociation
For flow cytometry of liver cells, single-cell suspensions were prepared from freshly excised mouse livers by mechanical trituration, after which the samples were passed through 70 mm steel mesh, and hepatocytes were isolated from the cell suspensions by centrifugation at 50 g for 5 min. The remaining cells were used for flow cytometry. For flow sorting of liver cells, livers were extracted and minced, and hepatocytes were isolated by mechanical trituration and centrifugation. The remaining liver tissue was digested with collagenase II (1 mg/ml) and DNase (100 µg/ml) in DMEM for 20 min at 37 °C. The cells were then filtered through 70-mm strainers to remove small fragments of undigested tissue for subsequent experiments.
Flow cytometry and sorting
Single-cell suspensions from mouse tissues were first stained with anti-mouse CD16/32 (BioLegend, #101320) to block the IgG Fc receptor, after which the cells were stained with surface fluorescent antibodies on ice for 30 min. The details of the antibodies used are listed in Table S3. Flow cytometry was performed on a Beckman CytoFLEX flow cytometer.
Isolation and culture of primary mouse hepatocytes
Mouse primary hepatocytes were isolated by liver perfusion medium using a 2-step retrograde procedure. Under terminal anesthesia, the mice underwent laparotomy, the inferior vena cava was then cannulated, and the superior vena cava was clamped to achieve retroperfusion of the liver using the portal vein as an outlet. The liver was perfused sequentially with buffer A (HBSS + 0.2 mg/ml EDTA + 1 mg/ml glucose) and then buffer B (HBSS + 0.75 mg/ml collagenase IV + 0.02 mg/ml DNase + 1 mg/ml glucose). After perfusion, the liver capsule was removed, and the liver was gently swirled in PBS to yield a cell suspension. Hepatocytes were collected by three rounds of centrifugation (50 g for 3 min) and cultured in DMEM/F12 supplemented with 10% fetal bovine serum, 1× ITS (Sigma‒Aldrich), 40 ng/ml dexamethasone (Sigma‒Aldrich) and penicillin/streptomycin.
Neutrophil isolation
Human neutrophils were isolated from the peripheral blood of healthy volunteers by density gradient separation using Ficoll (Cytiva, 17544202) and centrifugation at 400 × g for 40 min at room temperature. To isolate neutrophils from bone marrow, bone marrow cells from 8- to 12-week-old BALB/c mice were harvested in PBS, and the extraction of neutrophils from bone marrow cells was performed using a Mouse Neutrophil Isolation Kit (Solarbio) according to the manufacturer’s instructions.
Two-chamber neutrophil migration assays
Neutrophil migration assays were performed using a Transwell migration assay. Briefly, 5×105 freshly isolated neutrophils in RPMI 1640 were added to the upper chamber, and rhLRG1 (20 µg/ml) was added to the lower chamber as a chemoattractant. The migrated cells in the lower chamber were counted after 4 h.
In vitro NET analysis
To assess NET formation, neutrophils (1×106 cells) were seeded on coverslips coated with poly-L-lysine in 24-well plates for 30 min before rhLRG1 (20 µg/ml), PAD4i and SB431542 were added. After 6 to 8 h at 37 °C, the neutrophils were fixed with 4% paraformaldehyde (PFA) for 10 min at room temperature, washed twice with PBS, blocked in PBS containing 2% BSA for 30 min, and then incubated with anti-H3cit (1:100, ab5103, Abcam) and anti-MPO (10 µg/ml, AF3667, R&D) antibodies in blocking buffer overnight at 4 °C. After three washes in PBS, the cells were incubated with fluorochrome-conjugated secondary antibodies for 1 h and then counterstained with DAPI. Observation and imaging were performed with a confocal microscope (Cell Observer; ZEISS, Germany).
Transwell migration assays
For DLD1 cells and HCT116 cells (5×104), Transwell migration assays were performed. DLD1 cells and HCT116 cells were plated in the upper wells for 24 h. After the cells adhered, the medium was replaced with 300 µl of serum-free conditioned medium supplemented with neutrophils (1×105), rhLRG1 (20 µg/ml) or DNase (0.25 mg/ml). Complete medium supplemented with 10% FBS was added to the bottom to induce chemotaxis. After 48 h, the cells on the upper surface of the membrane were removed, and the cells that crossed the membrane were fixed with 4% paraformaldehyde and stained with crystal violet. The number of penetrated cells was counted under a light microscope in three fields of view, and the average number of cells was calculated.
Cytokine array analyses
Serum from the two CRC models was obtained by centrifugation and stored at −80 °C for use in cytokine assays. This was performed using the Mouse Inflammation Array GS1 (Raybiotech, Peachtree Corners, GA, USA) following the instructions of the manufacturer (Wayen Biotechnologies Inc., Shanghai, China).
RNA-seq data analysis
Raw RNA-seq sequencing data (FASTQ format) were aligned to the mouse reference genome mm10 using HISAT2 (v2.1.0) with default parameters to ensure alignment accuracy. Exon-aligned uniquely mapped reads were quantified using featureCounts (Subread package v2.0.6) to generate the raw count matrix. Gene expression data were normalized to transcripts per million (TPM), followed by hierarchical clustering analysis and heatmap visualization using pheatmap (v1.0.12). For the public dataset GSE109480 (retrieved from the GEO database), its raw expression matrix underwent identical TPM normalization, and differential gene expression bar plots were generated using ggplot2 (v3.5.0).
Single-cell sequencing data analysis
Single-cell sequencing data were processed using the Seurat (v4.2.0) pipeline. Strict quality control was applied: low-quality cells with total UMI counts <1000 or detected genes <200 were filtered out, along with apoptotic or damaged cells whose mitochondrial gene content was >25%. Red blood cells with hemoglobin gene expression (e.g., Hba-a1 and Hbb-bt) >1% were excluded. To mitigate doublet interference, DoubletFinder (v2.0.3) was employed to predict doublets.
Differential analysis and pathway enrichment
Differentially expressed genes (DEGs) were identified using the Wilcoxon rank-sum test via Seurat’s FindMarkers function, focusing on neutrophil subsets whose expression was significantly upregulated in the Lrg1(+/+)Hep-PMN and Lrg1(+/+)Hep-Ctrl groups (adjusted p value < 0.05, Benjamini–Hochberg correction; log-fold change [logFC] >0.5). For these DEGs, Gene Ontology (GO) functional enrichment analysis was performed using clusterProfiler (v4.7.1), with Benjamini‒Hochberg-adjusted q values < 0.05 as the significance threshold. The top 20 enriched terms were ranked by enrichment factors. For the PMN-MDSC signature score, individual cells were scored using the AddModuleScore function, which calculates the average expression levels of selected genes at the single-cell level and then subtraction by the aggregated expression of control feature sets.
Pseudotime analysis
We used Monocle3 (v1.3.1) to analyze the pseudotime distribution in the scRNA-seq data and construct cell trajectories to identify state transitions within the neutrophil populations. During trajectory and pseudotime computation, cells with high differentiation potential predicted by CytoTRACE (v0.3.3) were selected as the root nodes.
Statistical analysis
Statistical tests were carried out using GraphPad Prism (v9.5.0). Unless otherwise stated, the experimental data are presented as the mean ± standard deviation (SD) of at least three biologically independent replicates. To compare the parametric data, a two-tailed unpaired Student’s t test was used to determine statistical significance. A p value less than 0.05 was considered to indicate statistical significance.