Tissue-specific expression of PKM1 and PKM2 and its clinical significance in PDAC
Given that the expression and subcellular localization of the PKM1 and PKM2 splice isoforms are highly context dependent and that their quantitatively different enzymatic activities contribute to distinct metabolic modes in tumors10,19,26, we first investigated the expression of PKM1 and PKM2 in primary human PDAC tumors. PKM2 mRNA and protein expression were upregulated in PDAC tumors (Fig. 1a left, c top), and both were associated with poor patient prognosis (Fig. 1a right, d). In contrast, both PKM1 mRNA and protein levels remained low (Fig. 1b left, c bottom) and were not correlated with prognosis (Fig. 1b right). Additionally, PKM2 was upregulated in PanIN lesions and neoplastic epithelium, compared with morphologically normal pancreas acinar cells, ductal cells, and regions of acinar-to-ductal metaplasia (ADM) (Fig. 1c; Supplementary Fig. S1a). Notably, morphologically normal pancreatic ductal cells had weak PKM2 expression, whereas acinar cells did not detectably express PKM2. PKM2 expression was upregulated during ADM progression (Supplementary Fig. S1a). Furthermore, PKM2 was highly expressed in fibroblasts, tertiary lymphoid structures (TLSs) (Supplementary Fig. S1b, green), and nerve cells (Supplementary Fig. S1b, red), whereas PKM1 was strongly expressed only in nerve cells (Supplementary Fig. S1b, red) and weakly expressed in vascular smooth muscle cells (Fig. 1c, black).
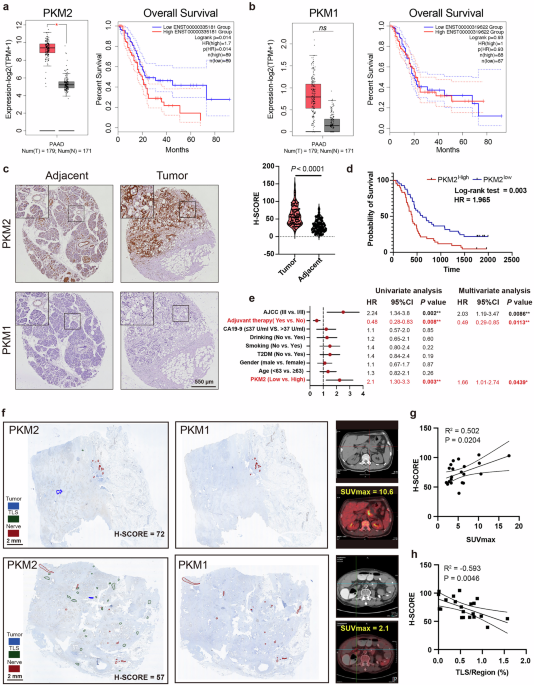
Fig. 1: Expression and localization of PKM1/2 in the PDAC TME and clinical significance.The alternative text for this image may have been generated using AI.
a, b mRNA levels of PKM1 and PKM2 and Kaplan–Meier survival analysis of patients with PDAC from The Cancer Genome Atlas (TCGA). Group cutoffs were set at the median mRNA expression of PKM2 (89 patients per group) or PKM1 (High = 88, Low = 87). c Representative immunohistochemical staining images with anti-PKM2 and anti-PKM1 antibodies. Violin plot showing the PKM2 IHC intensity score. Scale bar, 550 μm. Images are representative of 83 PDAC samples. d Overall survival based on the PKM2 IHC intensity score (High = 41, Low = 41). e Univariate and multivariate analyses of the OS of PDAC patients (n = 82 biological replicates). The dots on the forest plot represent the HRs of the Cox proportional hazards model, and the error bars are the two-sided 95% confidence intervals. *P < 0.05, **P < 0.01. Independent variables with P < 0.05 in the univariate analysis were included in the multivariate analysis. f Representative IHC staining image with anti-PKM2 and anti-PKM1 antibodies in patients with a high SUVmax (left) and a low SUVmax (right). The corresponding CT and PET images are shown on the right. Scale bar, 2 mm. Structures were manually annotated with Qupath: tumor (blue), TLS (green) and nerve (red). PKM1 is highly expressed in nerve areas only, whereas PKM2 is expressed ubiquitously. g, h Correlations between PKM2 intensity and the SUVmax (g) and between PKM2 intensity and the area of tumor-associated TLSs (h) from the PET/CT images of PDAC patients (n = 20 biological replicates). Statistical analysis: unpaired two-sided t-test (c); log-rank Mantel–Cox test (a, b, d).
Owing to the undetectable levels of PKM1 in the neoplastic epithelium, we were unable to further evaluate its correlation with the prognosis of PDAC patients. However, we identified PKM2 as an independent prognostic factor for overall survival (OS) (hazard ratio (HR) = 2.1; 95% confidence interval (CI): 1.30–3.3; P = 0.003) and as an unfavorable factor for adjuvant therapy (HR = 0.48; 95% CI: 0.28–0.83; P = 0.008) (Fig. 1e). This correlation is consistent with a previous report that the modulation of alternative PKM splicing and elevated PKM2 expression promote gemcitabine resistance27.
18F-Fluorodeoxyglucose (18F-FDG) positron emission tomography/computed tomography (PET/CT) has been widely used to visualize the metabolic activity of tumors. We therefore examined PET/CT images from PDAC patients and the corresponding paraffin-embedded, immunohistochemistry (IHC) tumor tissue sections. PKM2 was extensively expressed in the neoplastic epithelium (blue), TLSs (green), and nerves (red), whereas PKM1 was strongly expressed only in nerves (red) (Fig. 1f). Additionally, the intensity of PKM2 expression was positively correlated with the maximum standardized uptake value (SUVmax) (Fig. 1g). By normalizing the PKM2-positive TLS area to the entire slide area, we found that the PKM2 intensity was negatively correlated with the TLS area in PDAC patient samples (Fig. 1h), indicating the potential benefit of modulating PKM splicing to improve the response to immunotherapy.
Single-cell analysis reveals that tumor cells and activated Tregs rely on PKM2
To investigate the cell types that depend on PKM2 and that could be targeted to inhibit PDAC progression using our ASO, we reanalyzed a well-annotated single-cell RNA sequencing (scRNA-seq) dataset from patients with untreated, resectable, non-metastatic PDAC (n = 4)28. On the basis of the above PKM1 and PKM2 immunohistochemical staining results, we excluded minor cell types with strong expression of PKM1 only and focused on the remaining cell types that exhibited strong PKM2 expression (Fig. 2a, b). We investigated the unbiased most differentially expressed genes (DEGs) in addition to PKM2 expression (Fig. 2c). Uniform manifold approximation and projection (UMAP) analysis revealed that tumor cells formed two distinct subclusters that we annotated as basal-like (34.6%, 675/1952) and classical (65.4%, 1277/1952) PDAC cells, as previously described (Fig. 2c; Supplementary Fig. S2a)29. Compared with classical PDAC cells, basal-like PDAC cells exhibited higher expression of PKM (log2FC = 0.5, adj. P = 0.03) as well as other glycolysis hub genes, such as LDHA (log2FC = 0.63, adj. P = 2.85E-09), ENO1 (log2FC = 1.03, adj. P = 4.63E−24) and SLC2A3, which encodes the glucose transporter GLUT3 (log2FC = 1.5, adj. P = 0.001). Basal-like PDAC cells were also enriched in the epithelial–mesenchymal transition (EMT) pathway, hypoxia, and glycolysis, whereas classical PDAC cells were not significantly enriched in glycolysis or angiogenesis (Fig. 2d). Similarly, cancer-associated fibroblast Cluster 2 (CAF2) (no significant difference in PKM expression), macrophage Cluster 3 (mφ3) (log2FC = 0.4, adj. P = 7.28E−88), and tumor-associated neutrophil Cluster 1 (TAN1) (log2FC = 1.5, adj. P = 1.02E−43) also exhibited highly enriched glycolysis (Supplementary Fig. S2a).
Fig. 2: Broad screen for potential cells in the PDAC TME that may be targets of PKM-ASO.The alternative text for this image may have been generated using AI.
a Schematic and rationale of using scRNA-seq and scST to screen and show the localization of potential TME cell types that may be targeted by PKM-ASO. Well-annotated scRNA-seq data (n = 4 PDAC patients; biological replicates) from our previous studies were used3,28. The PKM1/PKM2 isoforms cannot be directly distinguished by scRNA-seq because of sparse coverage and because the only sequence differences between these isoforms map to exons 9 and 10, respectively. After removing nerve cells and Schwann cells (marked by CDH19, PLP1, and SOX10), which showed strong constitutive expression of PKM1 in our IHC data, PKM expression in the remaining cells in scRNA-seq and scST should be regarded as PKM2 rather than PKM1, providing the opportunity to identify potential cell types that can be targeted by PKM-ASO. b Representative IHC staining image with anti-PKM2, anti-PKM1, anti-PGP9.5, anti-S100β, anti-p75NRT, and anti-αSMA antibodies from serial sections of the TMA. Scale bar, 200 μm (bottom), 500 μm (top). PGP9.5 and S100β are nerve markers, p75NRT is a Schwann cell marker, and αSMA is a matrix marker. Images are representative of 83 PDAC samples with similar strong and constitutive PKM1 expression in nerves and ubiquitous PKM2 expression. c PKM2 expression across different cell types, annotated on the basis of the expression of marker genes. d GO analysis showing the different metabolic modes in basal-like and classical tumor cells. e UMAP plot showing 11 subclusters of T cells (top) and pseudotime analysis (bottom) of CD4-CCR7, CD4-FOXP3/CTLA-4, and CD4-FOXP3/BATF cells to show the dynamic differentiation and activation of Tregs. CD4-CCR7 was set as the root for the pseudotime analysis. f Multi-IF staining of anti-PKM2, anti-BATF, anti-CD4, and DAPI. CD4-FOXP3/BATF cells surround the tumor area. Activated Tregs (arrows) co-express BATF, PKM2, and CD4 and surround PDAC cells. The image on the right is a 2.5-fold magnified view of the activated Tregs indicated by the red arrow. Scale bar, 0.1 mm. *P < 0.05, **P < 0.01, ***P < 0.001. The Benjamini‒Hochberg method was used to correct for multiple hypothesis testing (d).
Notably, compared with naive CD4+ T cells (which express CD4/CCR7), terminally differentiated Tregs (which express BATF/TNFRSF4/CTLA430) expressed much higher levels of PKM (Fig. 2c, e; Supplementary Fig. S2b). Activated Tregs are an immunosuppressive subset of CD4+ T cells that can promote tumor progression by impairing an effective antitumor immune response30. We performed splicing analysis of published RNA sequencing (RNA-seq) data6 for nTconvs (naive conventional T cells, CD45RA+CD25–CD4+, corresponding to naive CD4+ T cells) and eTregs (highly suppressive effector Tregs, CD45RA–CD25highFOXP3highCD4+, corresponding to activated Tregs) to determine their PK isoform expression patterns. Mutually exclusive splicing analysis of PKM revealed that the average exon 9 inclusion values were 18% and 9% in nTconvs and eTregs, respectively (Supplementary Fig. S2c, d). Transcript counts revealed that both nTconvs and eTregs preferentially expressed PKM2 rather than PKM1 (Supplementary Fig. S2e). This preference may be related to the need for immune cells to rapidly respond to external stimuli, as glycolysis promotes rapid ATP production and immune cell proliferation, supplies metabolic intermediates, and regulates cellular functions to support immune responses31. PKM2 expression remained elevated during Treg differentiation, which is consistent with our scRNA-seq data analysis (Fig. 2c). Additionally, the expression of activated Treg markers, such as FOXP3, BATF, TNFRSF4, and two immune checkpoint genes, CTLA4 and PD1, was markedly elevated in eTregs in this bulk RNA-seq dataset (Supplementary Fig. S2f).
Multiple immunofluorescence (IF) staining assays revealed that activated Tregs surrounded the tumor epithelium (Fig. 2f). In addition, our data suggested that tumor cells attracted activated Tregs via the receptor–ligand pairs C3-C3AR1, APP-CD74, LPAR2-ADGRE5, CD55-ADGRE5, CD47-SIRPG, and PPIA-BSG (Supplementary Fig. S3a). Interaction between complement C3 and C3AR1 was observed only between basal-like cells and activated Tregs, but not between classical cells and activated cells, and was associated with poor prognosis in the TCGA-PAAD cohort (Supplementary Fig. S3,c). Treatment of BxPC-3 cells, a typical basal-like PDAC cell line, with a PKM splice-switching ASO, ASO1-TMO (Supplementary Table S2), induced significant splice switching, followed by slight downregulation of the expression of basal-like markers and C3 (Supplementary Fig. S3,b, d). Thus, our results demonstrate that tumor cells and the TME, especially basal-like PDAC cells and activated Tregs, are potential targets of our PKM-ASO.
Microarray-based spatial transcriptomics reveals that activated Tregs that surround basal-like tumor cells are highly glycolytic and have undergone EMT
We identified an intimate connection between basal-like PDAC cells and activated Tregs in the scRNA-seq data, but the localization of these two cell types in the immune microenvironment is unclear. To better understand the cross-talk between tumor cells and Tregs on the basis of their spatial distribution within tumors, we first investigated the functional differences among CD4-CCR7, CD4-FOXP3/CTLA-4, and CD4-FOXP3/BATF cells by performing Gene Ontology (GO) enrichment analysis using the scRNA-seq data. CD4-CCR7 cells were enriched in pathways related to the negative regulation of transcription, translation, and immune response, which is consistent with their immature phenotype and potential to differentiate (Fig. 3a, left). CD4-FOXP3/CTLA-4 cells were enriched in pathways associated with negative regulation of T-cell proliferation, inflammatory response to antigenic stimulus, and positive NF-κB transcription factor activity4, which reflected Treg maturation after stimulation (Fig. 3a, middle). Finally, CD4-FOXP3/BATF cells were enriched in pathways related to the inflammatory response, negative regulation of the T-cell response, response to hypoxia, and glycolysis, which reflect the adaptation of activated Tregs to the hypoxic TME and their reliance on aerobic respiration to maintain their functions, such as migration4 and immunosuppression5 (Fig. 3a, right). Notably, CD4-FOXP3/BATF cells highly expressed PKM and multiple immune checkpoint genes, such as TNFRSF4, HAVCR2, and CD274 (Fig. 3a, right).
Fig. 3: Microarray-based spatial transcriptomics reveals that basal-like tumor cells are surrounded by CD4-FOXP3/BATF cells.The alternative text for this image may have been generated using AI.
a GO analysis of the top 50 genes in each cell cluster. A P value < 0.05 was used for further visualization. Hypoxia and glycolysis pathways are enriched in CD4-FOXP3/BATF cells. b Schematic diagram of the scST analysis process. BayesSpace was used for spatial clustering, and the set of TOP50 genes in each cluster was input into EnrichR for GO analysis and visualization via a heatmap and UMAP. Cell2location was used for cell-type deconvolution on the basis of our scRNA-seq analysis. Then, CD4-CCR7, CD4-FOXP3/CTLA-4, and CD4-FOXP3/BATF cells were directly mapped onto the scST. We followed the previous annotation for the nerve, epithelial, stroma, and undefined regions in the scST2. Finally, UCell was used for signature scoring and visualization in UMAP as well as scST. Hallmark gene signatures were obtained from MSigDB. c Based on Cell2location, CD4-CCR7, CD4-FOXP3/CTLA-4 and CD4-FOXP3/BATF cells were mapped onto UMAP produced by BayesSpace. d GO analysis of each cluster shown in UMAP (top); violin plots show the frequency of CD4-CCR7, CD4-FOXP3/CTLA-4, and CD4-FOXP3/BATF cells in each GO cluster (bottom). e Spatial mapping of CD4-CCR7, CD4-FOXP3/CTLA-4, CD4-FOXP3/BATF, and nerve cells in each section (n = 4 biological replicates). First, UCell was used to calculate the BASAL-SCORE for epithelial cells, which were subsequently mapped onto sections to determine the spatial proximity between basal-like cells and Tregs. The PDAC basal-like tumor signature was obtained from a previous work29. f Frequency of CD4-FOXP3/BATF cells localized in high/low signature score areas (n = 4 biological replicates). The signature was obtained from MSigDB, and UCell was used for evaluation. Statistical analysis: unpaired two-sided t-test (f); Benjamini‒Hochberg method to correct for multiple hypothesis testing (a).
We next characterized how Tregs at different stages interact with the neighboring TME using previous spatial transcriptomics annotation and the ST analysis pipeline (Fig. 3b). After clustering by BayesSpace (Supplementary Fig. S4a, b), Cell2location revealed that CD4-CCR7, CD4-FOXP3/CTLA-4, and CD4-FOXP3/BATF cells mapped to different regions within the tumor (Fig. 3c). Activated Tregs utilize glycolysis to migrate into inflamed tissues4. Activated Tregs (CD4-FOXP3/BATF) were linked to hypoxia, cell mobility, and glucose homeostasis (Fig. 3d). In contrast, compared with activated Tregs, CD4-CCR7 cells were highly enriched in lipid catabolic processes, reflecting their different metabolism; whereas CD4-FOXP3/CTLA-4 cells appeared to be in an intermediate transition stage from CD4-CCR7 to CD4-FOXP3/BATF cells (Fig. 3d). After mapping nerve cells (which express PKM1), the basal-like score for each epithelial area, and Tregs, we found that activated Tregs were most likely to surround epithelial areas with high basal-like scores, especially in Sections 1 and 4. In contrast, the localization of CD4-CCR7 and CD4-FOXP3/CTLA-4 cells was more random and generally in the epithelial areas with low basal-like scores (Fig. 3e).
To assess whether the function of Tregs was reflected in their localization, all spots were first scored for glycolysis, fatty acid, hypoxia, angiogenesis, EMT, and the TGF-β pathway signature, with a 50th percentile cutoff used to determine low and high regions (Supplementary Fig. S4d). CD4-FOXP3/BATF cells were more frequently present in regions with high scores for glycolysis, hypoxia, angiogenesis, EMT, and TGF-β and less frequently in fatty acid metabolism of high-scoring regions, which is highly consistent with the presumed mechanism of action of our PKM-ASO (Fig. 3f). In contrast, CD4-FOXP3/CTLA-4 cells were preferentially located in regions with low hypoxia and EMT. Neither CD4-CCR7 nor CD4-FOXP3/CTLA-4 cells showed a metabolic preference (Supplementary Fig. S4c). Thus, blocking the mutual attraction between basal-like PDAC cells and activated Tregs could be a potential therapeutic strategy.
Basal-like PDAC cells are more sensitive to PKM-ASO than classical PDAC cells
The metabolic differences we observed between basal-like and classical PDAC cells suggested that these cells might respond differently to PKM-ASO (Fig. 2d). Although the expression of PKM2, PKM1, and the PKM2/PKM1 ratio did not differ between basal-like and classical cell lines, we detected differences in the expression of splicing regulators between these two subtypes (Supplementary Fig. S5a–e). ASO1-TMO effectively inhibited the proliferation of basal-like cell lines but had less of an effect on the proliferation of classical cell lines (Fig. 4a; Supplementary Fig. S5,d). To investigate the extent of splice switching in each subtype of PDAC cells, we transfected representative PDAC cell lines with varying concentrations of ASO1-TMO and analyzed the extracted RNA by radioactive RT-PCR. As expected, ASO1-TMO simultaneously increased the expression of PKM1 mRNA, decreased that of PKM2 mRNA, and increased that of PKMds mRNA, an isoform that is subject to nonsense-mediated mRNA decay (NMD), because of the presence of a premature termination codon in exon 11 when both exons 9 and 10 are skipped32 (Fig. 4b, c). Notably, splice switching occurred at a lower ASO concentration in BxPC-3 cells (EC50 = 8.55 nmol/L, basal score = 3) than in MIA PaCa-2 cells (EC50 = 23.44 nmol/L, basal score = –0.8). Consistent with the RNA data, PKM splice switching led to an increase in PKM1 and a decrease in PKM2 protein levels without changing total PKM expression in both MIA PaCa-2 and BxPC-3 cells (Fig. 4d). Collectively, these results confirm that our ASO1-TMO induces PKM splice switching and suggest that, compared with classical PDAC cells, basal-like PDAC cells are more sensitive to ASO1-TMO.
Fig. 4: Transfection of ASO1-TMO induces PKM splice switching in PDAC cells.The alternative text for this image may have been generated using AI.
a IC50 of ASO1-TMO. Three classical PDAC cell lines (SUIT-2, PANC-1, and AsPC-1), two basal-like PDAC cell lines (Hs 766 T and BxPC-3), and the MIA PaCa-2 cell line were used for IC50 determination. Cells were transfected with Lipofectamine and varying concentrations (0, 25, 50, 100, and 200 nM) of ASO1-TMO on Day 0, 103 cells were plated on a 96-well plate on Day 1, and cell viability was monitored by a colorimetric assay on Day 3. b Radioactive RT-PCR showing the extent of PKM splice switching after MIA PaCa-2 cells were transfected with the indicated concentrations of ASO1-TMO for 2 days (left). ImageJ was used to quantify the expression of the PKM2, PKM1, and PKMds isoforms (right). c Radioactive RT-PCR results showing the extent of PKM splice switching after BxPC-3 cells were transfected with various concentrations of ASO1-TMO as described in b for 2 days (left). Isoform quantification in b (right). d Western blot analysis showing the extent of isoform switching after MIA PaCa-2 or BxPC-3 cells were transfected with 50 nM ASO for 3 days. NTC, no-treatment control. e Viability of MIA PaCa-2 cells at each time point. Cells were transfected with 50 nM ASO on Day 0, 0.5 × 103 cells were plated on a 96-well plate on Day 1, and the OD450 was monitored daily by a colorimetric assay. f PK activity was measured on Day 3 after transfection with 50 nM ASO. g Lactic acid levels were measured on Day 3 after transfection with 50 nM ASO. Statistical analysis: unpaired two-sided t test (a, b, c, f, g); two-way ANOVA (e).
To further validate the antitumor efficacy of ASO1-TMO, MIA PaCa-2 and BxPC-3 cells were transfected with 50 nmol/L ASO and they showed significantly reduced proliferation compared with cells transfected with the scrambled sequence control, SCR-TMO (Fig. 4e; Supplementary Fig. S5,f). PKM1 is a constitutively active tetramer, whereas PKM2 is allosterically regulated and exists as either a catalytically active tetramer or a low PK activity dimer and monomer10 (Supplementary Fig. S5,g). ASO1-TMO significantly reduced the expression of PKM2 monomers/dimers, which are essential for aerobic glycolysis in tumors. Considering that ASO1-TMO induced splice switching from PKM2 to PKM1 and therefore depleted the low-activity PKM2 monomer/dimer, we expected that PK activity would increase and that glycolysis would decrease following ASO1-TMO treatment. Consistent with our previous work22, total PK activity significantly increased after ASO1-TMO treatment in both MIA PaCa-2 and BxPC-3 cells (Fig. 4f), and the level of the terminal glycolysis metabolic product lactate significantly decreased by approximately 12-fold and 8-fold in MIA PaCa-2 and BxPC-3 cells, respectively (Fig. 4g). Interestingly, treatment with si-PKM2 (Supplementary Fig. S5,h) decreased BxPC-3 cell proliferation but only had a limited effect on MIA PaCa-2 cells (Supplementary Fig. S5,i). Additionally, overexpression of PKM1 (Supplementary Fig. S5j) did not affect the proliferation of either cell line (Supplementary Fig. S5k). We conclude that the inhibition of cell proliferation caused by ASO1-TMO was due to the downregulation of PKM2 rather than the upregulation of PKM1. Our observations above revealed that the proliferation of MIA PaCa-2 cells decreased only slightly upon si-PKM2 treatment (with one of two siRNAs; Supplementary Fig. S5i) but that these cells were sensitive to ASO1-TMO treatment (Fig. 4a, e), which might reflect the fact that PKM1 can also promote PKM2 tetramer formation19 (Supplementary Fig. S5g).
PKM-ASO inhibits MIA PaCa-2 cell growth in an immunodeficient mouse model
Before administering the ASO in vivo, we first evaluated whether ASO1-TMO delivered by free uptake (gymnosis) could induce PKM splice switching in the above cell lines. Consistent with the results of the above transfection assays, ASO1-TMO had a minor but significant splice-switching effect at approximately 20 µmol/L in both BxPC-3 and MIA PaCa-2 cells (Supplementary Fig. S6a, b). At this same ASO concentration, we also observed significantly slower proliferation and reduced G2/M cell cycle arrest in MIA PaCa-2 and BxPC-3 cells (Supplementary Fig. S6c, d) along with increased PK activity (Supplementary Fig. S6e) and decreased lactate production (Supplementary Fig. S6f). We then explored the in vivo efficacy of unformulated ASO1-TMO in an orthotopic xenograft model (Fig. 5a). To rule out potential off-target effects in vivo, we also used a second splice-switching ASO, ASO2-TMO (Supplementary Table S2), with a different PKM target sequence22, which showed efficacy comparable to that of ASO1-TMO (Fig. 5a; Supplementary Fig. S6g).
Fig. 5: ASO1-TMO and ASO2-TMO inhibit the growth of MIA PaCa-2 orthotopic xenografts.The alternative text for this image may have been generated using AI.
a Schematic diagram of the animal experiment. ASO1-TMO and ASO2-TMO target the human PKM exon 10 regions (red and blue, respectively) that we previously targeted with MOE ASOs60. ASO2-TMO targets an SRSF3 binding site60 that is conserved between humans and mice (yellow). b Whole-animal live imaging of luciferase-expressing MIA PaCa-2 cells transplanted into the pancreas. Luminescence images of the transplanted mice on the indicated days are shown with a color scale in photons/sec/cm2/steradian. A total of 14 mice were randomized to each treatment group. c Quantification of the luciferase signal in (b). Statistical analysis: two-way ANOVA. d Images of representative tumors from animals treated as indicated on Day 23. The duodenum, pancreas, tumor, and spleen are shown. Dashed lines delineate the tumor. Scale bar, 1 cm. e Quantification of tumor weight on Day 23 (after the duodenum, normal pancreas, and spleen were removed). f ASO1-TMO and ASO2-TMO induce PKM splice switching in pancreatic tumors, as determined by radioactive RT-PCR of RNA from pancreatic tumor samples from tumor-bearing mice on Day 23. g Western blot analysis of PKM1, PKM2, and Vinculin in orthotopic tumor samples from mice treated with SCR-TMO, ASO1-TMO, or ASO2-TMO. h Representative H&E and IHC images of a tumor, showing the expression of PKM1 and PKM2. i Representative IF staining images of tumor sections from mice treated with saline, SCR-TMO, ASO1-TMO, or ASO2-TMO showing the expression of PKM2 (green), the proliferation marker Ki67 (red), and nuclei stained with 4′,6-diamidino-2-phenylindole (DAPI; blue). Scale bar, 50 µm. Statistical analysis: unpaired two-sided t-test (e, f); two-way ANOVA (c).
We used MIA PaCa-2 cells for the in vivo orthotopic xenograft experiments, as ASO treatment markedly inhibited their proliferation in vitro. After surgery, the tumors were allowed to engraft for 3 days prior to ASO treatment. The saline control group showed active tumor growth throughout the experiment (Fig. 5b). In contrast to the strong tumor inhibition induced by PKM-ASO treatment that was previously reported in two HCC mouse models22, the proliferation of xenografted PDAC tumors was relatively slow during the first two weeks but increased rapidly during the third week of PKM-ASO treatment (Fig. 5b, c). Nevertheless, the luciferase signal was significantly weaker in both ASO1-TMO-treated and ASO2-TMO-treated mice than in ASO-SCR-treated or saline-treated mice at the end of the experiment (Day 21) (Fig. 5c). PKM-ASO treatment was not associated with changes in body weight in this tumor model (Supplementary Fig. S7a). Additionally, fewer tumors were isolated from the pancreas of ASO1-TMO-treated and ASO2-TMO-treated mice on Day 23 than from ASO-SCR-treated mice (Fig. 5d, e).
Using radioactive RT-PCR, we detected significant splice switching in PDAC tumors following ASO treatment (Fig. 5f; Supplementary Fig. S7b). We also analyzed PKM2 and PKM1 protein expression, but detected only slight downregulation of PKM2 expression and did not consistently observe isoform switching after ASO treatment (Fig. 5g; Supplementary Fig. S7c). Considering the rapid tumor growth in the third week, isoform switching might be masked by negative selection or the development of treatment resistance (Fig. 5c).
We next performed IHC using anti-PKM2-specific and anti-PKM1-specific antibodies to evaluate metabolic heterogeneity. The PKM2 signal was slightly weaker in tumors treated with PKM-ASO, with a slight increase in the PKM1 signal in a minority of cells (Fig. 5h, arrow). To further characterize the localization of PKM-ASO and its effect, we first verified by IF that the ASO could be effectively delivered to PDAC tumors in vivo. As expected, we detected the ASO in PDAC tumor cells and a decrease in PKM2 expression, as well as a decrease in the nuclear area, indicating that these cells underwent necrosis or apoptosis (Supplementary Fig. S7e). We then focused on the necrotic areas on the basis of H&E staining. PKM-ASO significantly decreased the intensity of PKM2 and Ki67 staining (a proliferation marker) but did not significantly increase the intensity of PKM1 staining (Fig. 5i; Supplementary Fig. S7f, g). We conclude that treatment with PKM-ASO inhibits PDAC tumor growth in this xenograft model.
To evaluate the potential off-target and side effects of PKM-ASO, we analyzed kidney, liver, duodenum, and muscle tissues. We chose these tissues for the following reasons: first, ASOs generally accumulate in the kidney and liver, which are major organs affected by off-target toxicity24; second, muscle tissue expresses PKM1 rather than PKM2, so any histological changes there would be due to off-target effects; and third, the duodenum was reported to express PKM210, making it relevant to evaluate histological changes and splice switching for assessing potential side effects (Supplementary Fig. S7h). A comprehensive histopathological assessment revealed no significant changes or toxic effects of PKM-ASO or ASO-SCR in the kidney (Supplementary Fig. S7h, k), duodenum, liver, or muscle (Supplementary Fig. S7h). PKM1 and PKM2 expression did not significantly change in the duodenum (Supplementary Fig. S7i). Furthermore, there was no significant splice switching observed in the kidney or duodenum (Supplementary Fig. S7j, l, m). Collectively, these results position PKM-ASO as an effective PKM splicing modulator with no signs of toxicity as a single agent in preclinical models of PDAC.
Synergy between PKM-ASO and anti-CTLA-4 treatment in an immunocompetent PDAC mouse model
Considering the limited treatment efficacy of single anti-cancer agents, we evaluated potential synergistic treatments on the basis of the above PET/CT (Fig. 1f), scRNA-seq (Fig. 2c), and scST (Fig. 2e) data. Because PKM-ASO is also taken up by non-tumor cells, e.g., in our previous study of HCC22, and because inhibition of glycolysis can sensitize tumors to immunotherapy7,8, it was logical to consider the immune microenvironment in PDAC. We tested three potent murine Pkm ASOs on the basis of our previous screen22 with locked nucleic acid (LNA) or 2′-O-methoxyethyl (MOE) modifications along with two thiomorpholino (TMO)-modified ASOs whose target sequences overlap potential splicing factor binding motifs (Supplementary Fig. S8a and Table S2). We identified MOE16 as the most potent of these ASOs, which induced significant Pkm splice switching in both transfection and free uptake experiments with murine KPC1412 tumor cells (Supplementary Fig. S8b, c). Moreover, MOE16 strongly inhibited KPC1412 tumor cell growth in vitro in both the transfection and free uptake experiments (Supplementary Fig. S8d, e). The target site for MOE16 in Pkm partially overlaps the corresponding binding site for ASO2-TMO in the PKM ortholog, with only one base difference between the H. sapiens and M. musculus sequences. We also designed two MOE/PS-modified ASO controls, neither of which elicited splice switching compared with the MOE16-treated group nor had any obvious effect on KPC1412 cell viability compared with the NTC group (Supplementary Fig. S8f). We selected mSCR2 as a scrambled sequence control ASO for further in vivo study.
To determine whether inducing Pkm splice switching can synergize with anti-CTLA-4 therapy and sensitize PDAC cells to ICB, we treated established orthotopic allograft PDAC tumors with an anti-CTLA-4 antibody alone or in combination with MOE16 (Fig. 6a). Consistent with the results of previous pre-clinical experiments33 and those of a clinical trial34, the tumors from the control animals did not respond to anti-CTLA-4 treatment (Fig. 6b, c, f; Supplementary Fig. S9a). The tumors from animals treated with MOE16 alone were slightly smaller, which is consistent with our results in the immunodeficient mouse model (Fig. 6b, c; Supplementary Fig. S9a). Compared with MOE16 + IgG and mSCR2 + IgG, combined treatment with MOE16 plus anti-CTLA-4 had a synergistic effect, significantly decreasing tumor weight (Fig. 6b, c; Supplementary Fig. S9a). Notably, 2/8 tumors were undetectable by visual inspection. Moreover, this combination therapy increased the response rate to anti-CTLA-4 treatment (anti-CTLA-4 + mSCR2 vs IgG + mSCR2 = 12.5%, MOE16 + anti-CTLA-4 vs MOE16 + IgG = 75%) (Fig. 6f) without obvious toxicity (Supplementary Fig. S9b, c). Surprisingly, MOE16 treatment resulted in a slight increase in body weight (Supplementary Fig. S9b). Systemic delivery of MOE16 induced splice switching in PDAC tumors, the duodenum and kidney and the corresponding protein isoform changes in PDAC tumors (Fig. 6d, e, h; Supplementary Fig. S9,d, e). The tumors exhibited decreased infiltration of FOXP3+ Tregs, decreased Pkm2 mRNA expression, and increased Pkm1 mRNA expression (Fig. 6g, h; Supplementary Fig. S9f, g). Anti-CTLA-4 monotherapy did not significantly affect Treg infiltration in PDAC tumors, but Pkm-ASO monotherapy or combination therapy effectively reduced FOXP3+ Treg infiltration (Fig. 6g). CD4+ T cells that accumulated ASOs in the cytosol had decreased PKM2 expression (white triangle), whereas CD4+ T cells without ASO accumulation in the cytosol had high PKM2 expression (red triangle) (Fig. 6i; Supplementary Fig. S9h, i).
Fig. 6: ASO-based Pkm splice switching increases the response to anti-CTLA-4 therapy in PDAC models.The alternative text for this image may have been generated using AI.
a Schematic diagram of the experiment in C57/BL-6J mice. MOE16 (see Supplementary Fig. S8a) targets a conserved SRSF3 binding site and was delivered i.p. with a weekly schedule of 5 consecutive injections followed by 2 days of rest for 2 weeks. One hundred micrograms of mouse IgG2b control antibody or anti-CTLA-4 antibody was delivered i.p. every 3 days. b Images of representative tumors on Day 15; scale bar, 1 cm. c Tumor weight/weight of the whole pancreas on Day 15 (n = 8 biological replicates per group). d PKM splice switching reflected at the protein level in the mSCR2 + IgG and MOE16 + IgG groups (red, Vinculin, 124 kDa; green, PKM1, PKM2, total PKM, 60 kDa). ImageJ was used to quantify the PKM1, PKM2, and total PKM protein levels. e Radioactive RT-PCR of Pkm splice switching in tumors 15 days after i.p. injection of ASO. Tumor #4 in Row 3 of Panel b was not included because of low RNA quality. ImageJ was used to quantify the expression of the Pkm1 and Pkm2 isoforms. f Response rates measured from two independent experiments. Response was defined as a > 50% reduction in tumor weight compared with that of control tumors (mSCR2 + IgG). N/A, not applicable. g PKM2 intensity (left, based on h) and FOXP3+ cell area (right, based on Supplementary Fig. S9f) in each treatment group. h H&E staining and IHC results showing Pkm splice switching 15 days after i.p. injection of MOE16 or mSCR2 ASO and the anti-CTLA-4 or IgG antibody. The black arrow shows the peripheral area of the tumor. Scale bars, 500 μm (inset). i H&E staining, IHC, and IF staining results showing a pancreatic tumor in remission (tumor #4 in the 4th row of Panel b, following anti-CTLA4 and MOE16 ASO treatment). The white dashed line indicates tumor remnants. Scale bars, 250 and 100 μm (inset). Orange triangle: nerve. Green triangle: vessel. White triangle: ASO+CD4+PKM2– cells. Red triangle: ASO–CD4+PKM2+ cells. Yellow triangle: ASO–CD4+PKM2– cells. Zen was used to quantify the intensity of PKM2 and ASO in CD4+ T cells, and Pearson correlation analysis was performed. IHC experiments in h (including treatments with scrambled sequence ASO and IgG) and 6i were performed in parallel. Statistical analysis: unpaired two-sided t test (c, d, e, g); Pearson correlation analysis (i).