Patient characteristics
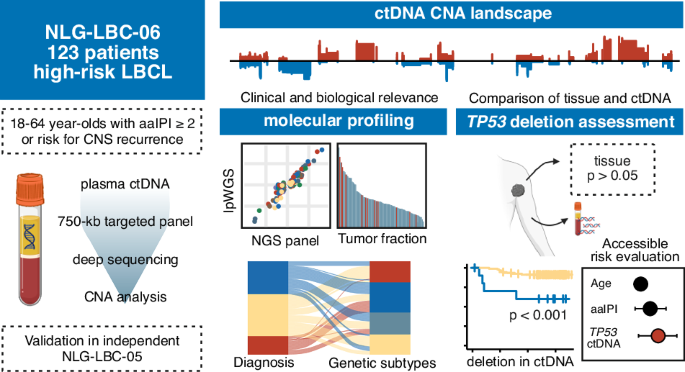
The patients (n = 123) included in the study were young (aged 18-64 years), had clinically high-risk (aaIPI ≥ 2 or site-specific risk factors for CNS recurrence) LBCL, and were treated uniformly in the Nordic phase II trial (Fig. 1A, Figure S1A, Table 1) with a dose-intensified immunochemotherapy [17]. The median follow-up time was 5 years at the time of the analysis. Plasma samples for ctDNA analyses were collected at baseline (BL), after two (CYC2) and/or after four cycles of therapy (CYC4), and at the end of treatment (EOT), and at follow-up (FU) (Fig. 1A, Figure S1A).
Fig. 1: Targeted ctDNA CNA landscape of LBC-06 patients.The alternative text for this image may have been generated using AI.
A Swimmer plot of the patients in the LBC-06 trial (n = 123). Purple lines denote cured patients, whereas red lines denote patients who died. Disease progression and death events are depicted as orange squares and red spheres. B Spearman’s correlation of overlapping segment means (log2, n = 322) detected with targeted panel (x-axis) and low-pass WGS (y-axis). Patients (n = 21) are colored differently. C Spearman’s correlation of overlapping CNA segment means (log2) in the ctDNA and segment means (log2) from matching FFPE tissues. Patients (n = 59) are colored differently. D Overall landscape of CNA proportions of n = 78 LBC-06 patients with n = 607 unique CNA segments across autosomal chromosomes. E Oncoprint of the most frequently aberrated regions. Gains and losses are colored with red and blue, respectively; copy number neutral regions are colored with yellow. Regions are annotated by cytobands, and the panel genes within these regions are marked in brackets. Every patient’s OS status, PFS status, diagnosis, DLBclass [39], and ctDNA burden are annotated on the oncoprint. lpWGS: low-pass whole genome sequencing, ctDNA: circulating tumor DNA, cfDNA: cell-free DNA, CNA: copy number aberration, OS: overall survival, PFS: progression free survival, GCB DLBCL: germinal center diffuse large B-cell lymphoma, NOS: not otherwise specified, THRLBCL: T-cell/histiocyte rich B-cell lymphoma, HGBL: high-grade B-cell lymphoma, FLG3b: follicular lymphoma grade 3B.
Targeted sequencing enables the detection of CNAs in LBCL
We applied a duplex adapter-informed 748-kilobase targeted gene panel and Illumina’s Dragen calling pipeline to 123 pretherapeutic plasma samples (Table S1, Materials and Methods). We hypothesized that the targeted panel, designed primarily for mutational profiling and MRD assessment, could also detect biologically relevant and clinically significant CNAs. First, to test our hypothesis, we compared the CNAs from targeted sequencing with the copy number landscape obtained by low-pass whole-genome sequencing (lpWGS; n = 21 patients; Fig. 1B), a more traditional method for profiling CNAs in ctDNA [34,35,36]. We found that the raw log2 copy number values produced with targeted sequencing correlated robustly with those from lpWGS (rho = 0.81, Fig. 1B), providing good sensitivity and excellent specificity (0.709 and 0.964; Figure S1B). In detail, 5% (n = 133) of the segment calls were discordant between lpWGS and panel data (Figure S1C). Additionally, we found that the raw log2 copy number values correlated well with those obtained using another publicly available analysis pipeline [34] (rho = 0.72, Figure S1D). This enabled us to examine the CNAs from FFPE lymphoma tissues as well. The comparison of CNA landscapes between matching ctDNA and tumor tissue showed a moderate correlation (rho = 0.5, n = 59, Fig. 1C), likely to be weakened by not only the spatial restrictions of a tissue biopsy [20, 21], but also the noise in the FFPE data [37] (Figure S1E).
Next, as the ctDNA tumor fraction is reported to affect the CNA detection across cancers [38], we were interested in determining the limit of detection (LOD) of our approach. Indeed, we detected that ctDNA VAF was associated with the number of detected CNAs (Figure S1F-G). To mitigate the effect of low ctDNA content on copy number calling, we restricted the analyses to samples with a mean VAF ≥ 0.015, corresponding to a tumor fraction of 0.03 [34]. This resulted in the assessment of 102 patients. The patient demographics of the 102 patients were similar to those of the whole cohort (Table 1). Notably, the 21 patients with a mean VAF < 0.015 had low disease burden (Figure S1H), which was reflected in excellent survival (Figure S1I-J). Finally, a 1:2 in-silico down-sampling experiment of 11 plasma samples revealed that reducing sequencing depth decreased detectable CNAs in the ctDNA (Figure S1K-L, Supplementary Materials and methods), highlighting that, in addition to VAF, adequate sequencing coverage is important for detecting targeted CNAs in ctDNA.
Altogether, we sequenced plasma cfDNA from 123 LBCL patients and discovered that a targeted panel can be applied to analyze CNAs in ctDNA. Moreover, considering the effect of ctDNA content, we restricted the analyses to patients with a mean VAF ≥ 0.015, resulting in CNA assessment in 102 high-risk LBCL patients.
CNAs in the ctDNA reveal biological heterogeneity in LBCLs
Next, we explored the CNA landscape (Fig. 1D, Table S2). Overall, the distribution of CNAs among patients was heterogeneous (Fig. 1D-1E), with the maximum number of individual CNAs captured by our targeted panel being 32 (mean 8). Out of 102 samples assessed, 78 (76%) had detectable CNAs, while 24 (24%) had no CNA calls (Fig. 1E, Table S2). The most recurrent CNAs in our data affected common aberrant genomic regions in DLBCL [6, 8, 12], including gains of 2p16, 3q27, and 12q13, which encompass the genes REL, BCL6, and KMT2D, respectively, and losses of 9p21 and 6p22, which affect CDKN2A, CDKN2B, and HIST1 genes (Fig. 1E). When we combined CNAs with variant data, we found that specific mutations co-occurred with either copy number gains or losses (Figures S2A-B). For instance, losses co-occurred with coding mutations in genes such as TP53, B2M, and HIST1H1E (Figure S2B), suggesting a bi-allelic inactivation of these genes.
To further examine molecular differences between LBCL patients, we investigated the CNAs by diagnostic subtypes. We detected subtype-specific CNAs between germinal center B-cell (GCB) and non-GCB DLBCL (Fig. 2A), prompting us to study whether these subtypes could be further characterized by minimally invasive genomic profiling. The assembly of somatic coding mutations, CNAs, and translocation profiles in ctDNA enabled us to implement the DLBclass [39] and LymphGen [6, 7] molecular clusters, both of which revealed genetic heterogeneity within the diagnostic LBCL subtypes (Fig. 2B, Figure S3A). Notably, we detected BCL2 and BCL6 translocations in ctDNA from multiple patients for whom FISH analysis was not available at diagnosis (Figure S3B). Apart from the discrepancy in the “Other” LymphGen subgroup caused by different confidence value thresholds, the two clustering methods mostly agreed on the genetic subtypes (Figure S3C), and the clustering confidence was comparable regardless of tumor fraction in the ctDNA (Figure S3D, cutoff appointed as per Chapuy et al. [39], Table S3).
Fig. 2: Subtype-specific heterogeneity detected by ctDNA CNAs.The alternative text for this image may have been generated using AI.
A Significantly different CNAs between GCB DLBCL (n = 36) and non-GCB DLBCLs (n = 46) in the discovery cohort. The frequency of CNAs per subtype is depicted in y-axis. P-value significance levels are marked on top, and genes are annotated at the bottom of the plot. B The diagnostic subtypes of the patients and their matched DLBclass subgroups [39]. DLBclass subgroups were analyzed from SNV, translocation, and CNA data. C, D Overall survival of study cohort and validation cohort together stratified by (C) DLBclass subtypes and (D) LymphGen subtypes. GCB DLBCL: germinal center diffuse large B-cell lymphoma, NOS: not otherwise specified, THRLBCL: T-cell/histiocyte rich B-cell lymphoma, HGBL: high-grade B-cell lymphoma, FLG3b: follicular lymphoma grade 3B; FISH: fluorescent in-situ hybridization, ctDNA: circulating tumor DNA, N/A: not available, OS: overall survival, mo: months.
To increase the statistical power of potentially prognostic molecular subtypes (Fig. 2B), we joined the molecular clustering data of the study cohort with data from an independent validation cohort (86 patients from the NLG-LBC-05 trial [40], who were treated with early high-dose methotrexate and dose-intensive immunochemotherapy, Supplementary Materials and Methods, Table S4). We found that the patients in the CNA-driven C2/A53 cluster had the worst OS and PFS (Fig. 2C-D, Figure S4A-B) and the highest tumor fraction in ctDNA (Figure S4C-D). Furthermore, the C2/A53 cluster remained an independent predictor of PFS after adjustment for ctDNA burden (Figure S4E), suggesting that although ctDNA abundance affects CNA calling, the clustering and prognostic effect are driven by the underlying molecular aberrations. When we conducted the DLBclass analysis only with coding mutations and structural variants, there was no difference in patient outcomes between the molecular subgroups (Figure S4F-G). These results, together with the requirement for CNA data for A53 inclusion in LymphGen classification, underscore the importance of CNA profiling for accurate subtype-specific survival prediction.
Taken together, we found CNAs in the ctDNA that uncover biological heterogeneity among established subtypes and enable detailed genetic subclassification of LBCL from LB. These results highlight the strong complementary potential of ctDNA for subtype characterization of LBCL patients when diagnostic tumor tissue is limited.
High tumor fraction and multiple CNAs detect high-risk patients
The survival association identified by genetic subgroups cued us to further explore associations between copy number landscapes and clinical characteristics. High copy number-derived tumor fraction in the LB has been shown to correlate with more aggressive DLBCL [41], and, likewise, we found that tumor fraction was associated with worse OS, high VAF in the ctDNA and aaIPI, but not with age (Fig. 3A, B) and was similar between panel-based and lpWGS data (n = 26, Figure S5A). We were able to validate the results in the validation cohort (Fig. 3C, Figures S5B-D). The estimates of tumor fractions were comparable between the discovery and validation cohorts (mean 0.26, range 0.02-0.75, and mean 0.30, range 0-0.86, respectively, Fig. 3A, Figure S5B), suggesting that high-risk LBCL patients have similar tumor fractions, enabling consistent risk assessment in the LB.
Fig. 3: Characterization of high-risk LBCL patients using CNA landscapes.The alternative text for this image may have been generated using AI.
A Waterfall plot of tumor fractions in LBC-06 patients. Patients who died are colored orange, whereas patients who survived are colored blue. The gray dashed line represents a tumor fraction of 0.03, corresponding to the assay’s detection limit. Mean ctDNA VAF, age, and aaIPI are annotated at the bottom of the plot for each patient, and their correlation with tumor fraction is marked on the right. B Overall survival of patients stratified by tumor fraction. An optimal cutoff for tumor fraction (24%) was used as a threshold between patients with high and low tumor fractions. C Overall survival of validation cohort (LBC-05) stratified by tumor fraction. The discovery cohort’s cutoff for high tumor fraction (24%) was used to separate patients into high and low tumor fraction groups. D, E Univariate Cox regression analysis for OS of recurrent gains (D) and losses (E). The hazard ratio is depicted on the x-axis, and the -log10 p-value is on the y-axis. Aberrations in genes that reached statistical significance to OS are colored red (D) and blue (E). See also Table S5. ctDNA: circulating tumor DNA, VAF: variant allele frequency, OS: overall survival, PFS: progression-free survival, mo: months, aaIPI: age-adjusted International Prognostic Index, HR: hazard ratio, CI: confidence interval.
Next, we investigated whether individual CNAs could identify patients at high risk. By systematically applying the Cox regression model on recurrently altered regions ( ≥ 5 patients affected), we found that several CNAs, including BCL6 gain and TP53 loss, were associated with poor survival (Fig. 3D-E, Figures S5E-F, Table S5). The prognostic CNAs did not associate with clinical high-risk disease characteristics (Figure S5G), revealing heterogeneity within the clinical metrics used for patient stratification. Furthermore, we found that the number of prognostic CNAs cumulatively reflected survival (Figure S6A-B) and the C2 molecular subtype (Figure S6C). Harboring one or more of these high-risk CNAs remained as an independent prognostic factor for OS and PFS in multivariable analysis with age, aaIPI, and ctDNA concentration (Figure S6D-E).
Altogether, we observed similar tumor fraction landscapes in the LB and found that high tumor fraction is associated with worse survival in two independent patient cohorts. Additionally, we identified several prognostic CNAs that revealed clinical heterogeneity and found that harboring one or more of these aberrations was associated with worse survival cumulatively.
TP53 loss reveals a clinically relevant group of patients
Our data had so far highlighted the loss of TP53/17p in both subtype clustering and as a prognostic factor. To further explore the translational potential of TP53 loss, we examined it in the context of an already established marker used in risk stratification: a FISH-informed TP53 status. Overall, the proportions of TP53 statuses differed between the two assessment methods (chi-square test, p = 0.001, Fig. 4A). Upon closer examination, we found that several TP53/17p FISH-negative patients, indicating copy number neutrality, exhibited detectable TP53 loss in their ctDNA (Fig. 4B). The same result was obtained with another CNA segmentation pipeline (Figure S7A-S7B). When we investigated patients with negative FISH results but CNA loss in their ctDNA, we found that all but one harbored a coding SNV in TP53 (Fig. 4C). Overall, TP53 mutations in ctDNA were mostly pathogenic [42] (Figure S7C), and 46% of patients with a TP53 mutation had a TP53 loss (Fig. 4D), suggesting a common LOH event in lymphomagenesis. Furthermore, the ctDNA TP53 loss was associated with P53 positivity assessed by immunohistochemistry (Figure S7D). On the other hand, nine patients had TP53/17p FISH positivity despite undetected ctDNA loss (Fig. 4B). These patients had lower ctDNA concentration and tumor fraction than the patients with detectable TP53 CNA (Figure S7E-S7F).
Fig. 4: TP53 status reveals a clinically relevant subgroup of LBCL patients.The alternative text for this image may have been generated using AI.
A Comparison of TP53 status by two approaches: clinically used FISH from tumor tissue and ctDNA analysis. Chi-square test. Patient and plasma tube figures were created in BioRender.com, Arffman, M., 2025, https://BioRender.com/7xb3vjy. B Detailed differences in TP53 statuses between tumor tissue FISH and ctDNA analyses. TP53/17p FISH status in x-axis and copy number segment mean (log2) in y-axis. Patients with TP53 loss, TP53 gain, and copy number neutral TP53 in the CNA analysis are colored blue, orange, and yellow, respectively. C Non-silent TP53 variants in the ctDNA and the variant allele frequencies in all patients. Orange points depict variants from patients with negative FISH TP53/17p status but loss in ctDNA with ClinVar estimates for variant pathogenicity. Blue points depict TP53 variants from other patients. D Oncoprint of non-silent TP53 variants in all patients together with TP53 CNA and TP53/17p FISH statuses. Co-occurrence of the variant together with TP53 loss is depicted on the left (only ctDNA-informed TP53 loss: upper panel, both ctDNA- and FISH-informed TP53 losses: lower panel). E Survival analysis for OS of the patients stratified by ctDNA TP53 CNA status (n = 102). F Survival analysis for OS of all patients stratified by FISH TP53/17p status (n = 114). G Multivariable analysis for OS (n = 102): age, aaIPI, and ctDNA TP53 loss (n = 11). FISH: fluorescent in-situ hybridization, ctDNA: circulating tumor DNA, N/A: not available, VAF: variant allele frequency, CNA: copy number aberration, IHC: immunohistochemistry, OS: overall survival, aaIPI: age-adjusted International Prognostic Index, HR: hazard ratio, PFS: progression-free survival, mo: months, VUS: variant of uncertain significance.
After detecting a discrepancy between ctDNA and FISH-informed methods, we next evaluated the prognostic value of these assessments. Even though we did not detect TP53 losses in patients with small tumor fractions, ctDNA-assessed TP53 loss was associated with poor survival (Fig. 4E, Figure S7G), and, notably, was more prognostic than FISH-informed TP53/17p status both for OS (Fig. 4E-F) and PFS (Figure S7G-S7H). ctDNA-informed TP53 loss remained prognostic for OS in multivariable analysis with age and aaIPI (Fig. 4G, Table S6), whereas FISH-informed TP53/17p status did not (Figure S7I). Notably, ctDNA TP53 loss remained an independent predictor for OS even after adjusting for ctDNA burden or tumor purity (Figure S7I, Table S6), underlining that despite the positive correlation with tumor content in cfDNA (Figure S7E-S7F), minimally invasive detection of TP53 loss improved survival estimation. We observed that TP53 loss in ctDNA was also associated with worse PFS in the validation cohort (Figure S7J). Finally, although the co-occurrence of coding TP53 mutations and TP53 loss in ctDNA was associated with poor survival (Figure S7K), it did not improve survival assessment compared to CNA-based assessment alone. The results suggest that TP53 loss in ctDNA can serve as an independent marker to identify high-risk patients.
Altogether, these results demonstrate that LB-based TP53 assessment improves risk stratification over traditional a tissue-based method, enabling more accessible yet clinically relevant molecular profiling.
Copy number profiles and cancer cell fractions reveal structures of lymphoma progression
Finally, we were interested in studying ctDNA dynamics in sequential samples. Although most patients had no detectable CNAs in samples obtained during or after therapy due to diminished ctDNA levels, we wondered if the copy number landscape could inform about the mutational structure of non-responding lymphomas. Accordingly, while some patients with R/R LBCL showed a reduction in their CNAs during therapy (Patients #1-#2, Fig. 5A), others acquired new CNAs (Patients #3-#4, Fig. 5B). These results indicate that LBCLs adopt distinct mutational mechanisms in progression.
Fig. 5: Analysis of clonal population structures in R/R patients.The alternative text for this image may have been generated using AI.
A, B CNA landscapes of patients with less CNAs (Patient #1 and Patient #2, A) and more CNAs (Patient #3 and Patient #4, B) in the R/R plasma sample compared to pretreatment plasma sample. Selected genes are annotated below the landscape plots. C Patient #1 molecular dynamics: cancer cell fractions of distinct clones before treatment and at disease progression (left panel). Clones are marked with different colors, and selected genes in significantly expanding clones are annotated. ctDNA concentration at baseline, throughout treatment, and at disease progression (right panel). The gray dashed line at -1.5 denotes the threshold for MRD test positivity. D Patient #2 molecular dynamics: cancer cell fractions of distinct clones before treatment and at disease progression (left panel). Clones are colored with different colors, and selected genes in significantly expanding clones are annotated. ctDNA concentration at baseline, throughout treatment, and at disease progression (right panel). The gray dashed line at -1.5 denotes the threshold for MRD test positivity. E Patient #3 molecular dynamics: cancer cell fractions of distinct clones before treatment and at disease progression (left panel). Clones are marked with different colors, and selected genes are annotated. ctDNA concentration at baseline, throughout treatment, and at disease progression (right panel). The gray dashed line at -1.5 denotes the threshold for MRD positivity. F Patient #4 molecular dynamics: cancer cell fractions of distinct clones before treatment and at disease progression (left panel). Clones are marked with different colors, and selected genes in significantly expanding clones are annotated. ctDNA concentration at baseline, throughout treatment, and at disease progression (right panel). The gray dashed line at -1.5 denotes the threshold for MRD test positivity. GCB DLBCL: germinal center diffuse large B-cell lymphoma, CNA: copy number aberration, CCF: cancer cell fraction, R/R: relapsed/refractory, BL: baseline, CYC2: after 2 cycles, EOT: end of treatment, MRD: minimal residual disease, FU: follow-up, Mo: month.
To dynamically model cancer cell population structures across different time points [43], we combined CNA and SNV data from the ctDNA. Strikingly, duplex correction enabled highly detailed characterization of the cancer cell fraction (CCF) of the distinct clones (Figure S8A-B). Despite MRD negativity at the EOT (Patient #1), the main clones at pre-treatment were also frequently prominent at progression (Fig. 5C-F). However, we found shifts in CCFs between the two time points in all patients (Figure S8C). For example, a significant expansion of clones that harbored mutations in MYC and CD79B (Patient #1, Fig. 5C), PDCD1LG2 (Patient #2, Fig. 5D), and B2M, CD83, and IGLL5 (Patient #4, Fig. 5F) was detected (Figure S8C). Furthermore, emerging CNAs in patients #3 and #4 (Fig. 5B) that did not overlap with any SNVs suggest that there are additional clonal characteristics beyond SNV-dependent CCF estimates.
Collectively, these findings indicate that lymphoma evolution is a dynamic process, which can be revealed by joint analysis of CNAs and smaller variants in the LB. Although descriptive and mostly hypothesis-generating, these examples highlight the expanding opportunities for minimally invasive profiling in a sequential setting and suggest that, despite frequent fluctuations in CCFs, disease progression may often be driven by the founding clone.