Cohort description
For the discovery phase of this study, we utilized colon adenocarcinoma (COAD) samples obtained from TCGA10. The subsequent replication phase was carried out through a meta-analysis, incorporating data from studies on colon cancer (GSE131013, GSE42752, E-MTAB-7036 and GSE199057)11,12,13,14, rectal cancer (TCGA-READ and GSE39958)15 and colorectal cancer (GSE101764, GSE77954 and E-MTAB-3027)16,17,18. Patients were classified as EOCRC (<50 years) or LOCRC (≥70 years). This resulted in a discovery dataset containing 31 EOCRC and 100 LOCRC patients, whereas the replication cohort incorporated 83 EOCRC and 272 LOCRC patients. Population characteristics are presented in Table 1, with a detailed discovery cohort characterization in Supplementary Table 1.
Table 1 Study population characteristics
Exposome-related DNA methylation marker sets
The exposome’s association with EOCRC versus LOCRC was evaluated using 29 lifestyle and environmental factors, which we define here as the exposome, acknowledging its broader scope. The analyzed traits encompassed 11 lifestyle factors: the Alternative Healthy Eating Index (AHEI), alcohol consumption, birthweight, body mass index (BMI) (continuous variable in kg m−2), cannabis use, coffee consumption, education level, Mediterranean Diet Score (MDS), obesity (defined as ≥30 kg m−2), smoking habits and smoking inference model (smoking-Maas). Furthermore, we examined four air pollution particles: nitrogen dioxide (NO2), polychlorinated biphenyls (PCBs) and particulate matter (PM) < 2.5 µm in diameter (PM2.5) and between 2.5 µm and 10 µm (PM2.5–10). In addition, we included 14 pesticides encompassing 2,4-dichlorophenoxyacetic acid (2,4-D), atrazine, acetochlor, chlordane, dicamba, malathion, dichlorodiphenyltrichloroethane (DDT), heptachlor, lindane, glyphosate, mesotrione, metolachlor, picloram and toxaphene. For the marker selection, we selected for each trait significantly associated CpG sites (CpGs) from extensive epigenome-wide association studies (EWAS), employing various significance thresholds, namely genome-wide (GW: P < 1.2 × 10−7), P < 1.0 × 10−5 and false discovery rates (FDR) of <0.01, <0.05 and <0.1. Smoking-Maas represents the CpGs included in a previously developed smoking inference model and their published coefficients19; no additional selection threshold was applied in the current study.
Exposome-related MRSs
Using available EWAS summary statistics across the five marker selection thresholds, we identified 63 exposome CpG sets across 29 exposome traits and computed 63 weighted MRSs from DNA methylation beta-values adjusted for Horvath epigenetic age20. Although termed ‘risk scores,’ these MRS serve as proxies for exposome exposure, enabling the assessment of exposure prevalence in early- versus late-onset tumors rather than estimation of individual cancer risk. The number of CpGs and their weights can be found in Supplementary Tables 2 and 3.
The limited availability of exposome data in tumor samples hinders the evaluation of its impact on EOCRC and restricts direct validation of the constructed MRSs against measured exposures. To address this, we performed a comprehensive validation of the constructed MRSs across several datasets, with all detailed results summarized in Supplementary Table 5.
Validation was performed by reconstructing the MRS in a dataset with available exposure measurements and assessing concordance between MRS values and the corresponding recorded exposure. By leveraging TCGA data, we observed a higher alcohol MRS in liver cancer patients with alcoholic liver disease (ALD), and a separation of current and former smokers from never smokers with the smoking-Maas across several cancer types (Supplementary Table 4 and Extended Data Fig. 1). We further validated the use of MRSs as proxies for cannabis dependence, higher PM2.5 median exposure, and smokers (smoking-Maas) using publicly available datasets, including noncancer participants (Extended Data Fig. 1 and Supplementary Table 5).
Beyond the exposome traits included in our study, we examined additional traits using EWAS and publicly available datasets with DNA methylation and measured exposome data, including allergic asthma, major depressive disorder (MDD), polygenic risk scores (PRS) for depression, as well as prenatal exposures, including birthweight, gestational age, maternal education, maternal BMI and arsenic exposure. The constructed MRSs for these traits showed concordance with directly measured exposures, further strengthening the validity of MRSs as a proxy for exposure (Supplementary Table 5 and Extended Data Figs. 2–3).
Exposome-related methylation risk scores in EOCRC versus LOCRC
To elucidate the exposome’s impact on early-onset colon and rectal cancer cases, we compared the 63 MRSs between early-onset and late-onset (reference group) patients using multivariate logistic regression models. Due to sex disparities in CRC incidence21, we adjusted the regressions for sex when feasible. Figure 1 summarizes the results for all 29 exposome traits, analyzed across the five selection thresholds used in the original EWAS studies. In the discovery dataset, positive associations were observed for MRSs related to PCB, PM2.5, smoking-Maas, heptachlor, metolachlor, picloram and toxaphene. In contrast, negative associations were found for MRSs corresponding to BMI, education level, MDS, obesity, atrazine, malathion and mesotrione (Fig. 1 and Supplementary Table 6). We highlight the results for four lifestyle factors previously linked to colon and rectal cancers, including the MDS22 (Fig. 2a) and education level23 (Fig. 2b), which are considered protective factors, as well as smoking habits24 (Fig. 2c) and obesity status25 (Fig. 2d), which are recognized as risk factors. To examine the directionality of our findings, the heatmaps in the left panels of Fig. 2 show the methylation level distributions across CpGs featured in each of the four MRSs, along with their direction in the original EWAS and after sorting by the derived MRSs. The heatmaps show that increased MRS correlates with higher beta-values in CpGs with positive associations in the EWAS and with lower beta-values in CpGs with negative associations (see Extended Data Fig. 4 for a more detailed explanation). These results suggest that an elevated MRS reflects greater exposure levels in the original EWAS. Specifically, for patients with early-onset colon cancer, this suggests a lower MDS (P = 7.9 × 10−3; adjusted P (Padj.) = 3.3 × 10−2) (Fig. 2a), lower education levels (P = 4.0 × 10−3; Padj. = 2.2 × 10−2) (Fig. 2b), increased smoking (smoking-Maas) exposure (P = 1.1 × 10−3; Padj. = 8.9 × 10−3) (Fig. 2c), and lower obesity rates (P = 8.4 × 10−4; Padj. = 7.5 × 10−3) (Fig. 2d) in comparison to those with late-onset, as illustrated in the middle panels of Fig. 2. The association of lower obesity rates with early-onset cases was verified utilizing physical metrics from TCGA-COAD. Colon cancer patients with a BMI over 30 kg m−2, as measured in the clinic, were categorized as obese, resulting in 4 out of 24 early-onset and 18 out of 72 later-onset patients being marked as obese. This provides a relative risk (RR) of 0.67 (95% confidence interval (CI): 0.26–1.76) for obesity in early-onset colon cancer patients within the TCGA-COAD cohort (Extended Data Fig. 5), supporting the MRS results for obesity.
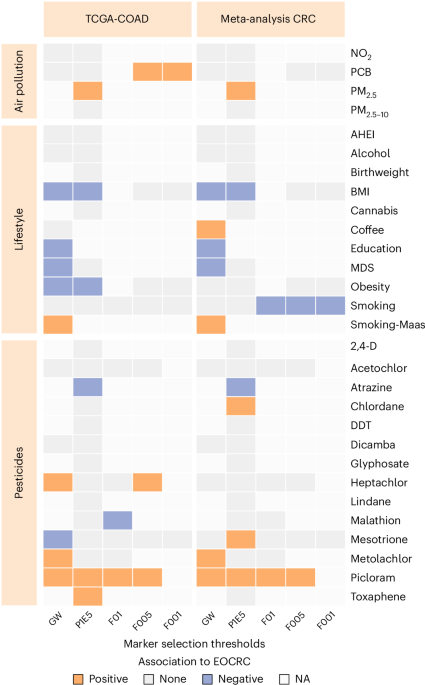
Fig. 1: Exposome-related methylation risk scores show differences between early- and late-onset CRC patients.The alternative text for this image may have been generated using AI.
Summary overview of the association between the 63 exposome-related MRSs across 29 exposome traits (y axis) and five selection thresholds (x axis, as applied in the original EWAS), comparing early-onset (age <50 years) to late-onset (age ≥70 years, reference) cancer using sex-adjusted logistic regression. The discovery phase (left panel) was conducted in TCGA-COAD (31 early-onset versus 100 late-onset colon cancer). The replication phase (right panel) shows meta-analysis results from nine cohorts (E-MTAB-3027, E-MTAB-7036, GSE199057, GSE131013, GSE42752, TCGA-READ, GSE39958, GSE101764 and GSE77954), including 83 EOCRC and 272 LOCRC patients. The selection thresholds (x axis) indicate the significance threshold in the original EWAS used for CpG selection, namely P < 1.2 × 10−7 (GW), P < 1.0 × 10−5 (P1E5) and FDRs of <0.1 (F01), <0.05 (F005) and <00.1 (F001). The presence of CpGs across the five selection thresholds varies, with unavailable data indicated in white (NA), and nonsignificant findings in light gray.
Fig. 2: Lifestyle factors in EOCRC versus LOCRC tumors.The alternative text for this image may have been generated using AI.
a–d, Lifestyle-related MRS differed between EOCRC and LOCRC, including non-Mediterranean dietary patterns MRS-GW (marker threshold P < 1.2 × 10−7) (a), lower education levels MRS-GW (b), higher smoking exposure (smoking-Maas) (c) and lower obesity rates MRS-P1E5 (marker threshold P < 1.0 × 10−5) (d) in patients with EOCRC. The heatmaps (left) display epigenetic age-adjusted DNA methylation beta-values in TCGA-COAD, ordered by MRS; the top color bar indicates CpG effect direction in the original EWAS (red: positive; blue: negative). The boxplots (middle) show MRS distribution in patients with EOCRC (N = 31) and LOCRC (N = 100) in TCGA-COAD, stratified by sex (orange: female; purple: male). The forest plots (right) present sex-adjusted logistic regression results for each replication dataset, their meta-analysis (Replication), and combined with TCGA-COAD (Discovery and Replication), for colon (blue), rectal (purple) and CRC (red). Squares: ORs; horizontal lines: 95% CIs. Boxplots show median (center line), interquartile range (IQR) (box) and whiskers (1.5× IQR) and outliers.
Extending our investigation, we conducted a meta-analysis across the nine replication datasets (Figs. 1 and 2 and Supplementary Table 7). This analysis corroborated the initial findings, notably the associations of a lower MDS (P = 1.5×10−2; Padj. = 4.6×10−2), lower educational levels (P = 2.11 × 10−5; Padj. = 3.59 × 10−4) and higher smoking exposure (P = 1.02 × 10−5; Padj. = 8.6 × 10−4) in EOCRC (Fig. 2, right panel). To ensure platform consistency, the meta-analysis was repeated using eight datasets generated with the 450K array, excluding the EPIC-based cohort GSE199057. This strengthened the association for lower MDS (P = 3.06 × 10−3; Padj. = 1.3 × 10−2), whereas educational level and smoking-Maas remained significant (Supplementary Table 8). Finally, despite the considerably reduced sample size, an exploratory tissue-specific meta-analysis of rectal-only (TCGA-READ and GSE39958) or colon-only cancer datasets (GSE131013, GSE42752, E-MTAB-7036 and GSE199057) showed a consistent trend in colon for education and smoking (Extended Data Fig. 6 and Supplementary Table 9).
Picloram-related MRSs
Our results highlight a new association between the MRSs for the herbicide picloram and EOCRC, in comparison to LOCRC cases, in both the discovery and meta-analysis (Fig. 1). We observed that a higher exposure level, as indicated by the original EWAS direction, is associated with an elevated MRS (Fig. 3a). This association highlights increased exposure to picloram among patients with early-onset colon cancer (P = 2.59 × 10−5; Padj. = 4.41 × 10−4) (Fig. 3b)—a finding consistently supported by our meta-analysis using all replication cohorts (P = 3.07 × 10−3; Padj. = 1.51 × 10−2; odds ratio (OR): 1.56 [95% CI: 1.16–2.09]) (Fig. 3c) as well as in the meta-analysis using only 450K array samples (P = 7.17 × 10−4; Padj. = 4.06 × 10−3; OR: 1.71 [95% CI: 1.25–2.33]).
Fig. 3: A higher picloram MRS is associated with EOCRC.The alternative text for this image may have been generated using AI.
The figure shows the distribution and validation of the picloram MRS employing the genome-wide CpG selection threshold (MRS-GW). a, Heatmap of epigenetic age-adjusted DNA methylation beta-values in TCGA-COAD, ordered by MRS; the top color bar indicates CpG effect direction in the original EWAS. b, Boxplot showing MRS-GW distribution in early- (N = 31) and late- (N = 100) onset TCGA-COAD patients, stratified by sex (orange, female; purple, male). c, Forest plot showing sex-adjusted logistic regression results in replication cohorts, their meta-analysis (Replication), and combined discovery and replication, including colon (blue), rectal (purple) and colorectal (red) cancer. d,e, The associations were further adjusted for other MRSs (31 EOCRC versus 100 LOCRC) (d) and tumor purity, subdivision (left/right), first-degree family with cancer diagnosis (yes/no), race (White/non-White) and MSI status (e). The baseline model (red) includes sex adjustment only; subset analyses show sex-adjusted (M1, orange) and fully adjusted models (M2, blue square). f, Spearman correlation between the 37-gene picloram-specific ssGSEA and the picloram MRS in TCGA-COAD (P = 2.127845 × 10−12); solid line: linear fit with the shaded area indicating the 95% CI. g,h, The violin plot shows the permutation results, confirming the robustness of CpG selection, patient categorization (g) and ssGSEA association (h). In the forest plots, squares represent ORs, and lines 95% CIs. Boxplots show median (center line), IQR (box), whiskers (1.5× IQR) and outliers.
The model presented was initially adjusted only for sex to facilitate external validation in independent datasets lacking measurements for additional variables. To evaluate the robustness of the association between picloram MRS-GW and age at onset, we applied additional adjustments in TCGA-COAD patients. First, we accounted for the influence of other MRSs included in the GW marker selection threshold (Fig. 3d). We observed only minor variations across most adjustments, and none resulted in the loss of the significant association between picloram MRS-GW and age at onset (Fig. 3d). Second, we explored the distribution of the picloram MRS-GW between male and female participants (P = 0.0024) and between microsatellite stable (MSS) and instable (MSI) tumors (P = 0.01), tumor purity high (≥0.7) versus low (<0.7) (P = 5.3 × 10−15), and across the consensus molecular subtypes (Extended Data Fig. 7). The observed significant differences highlight the need for further adjustment when testing the association between picloram MRS-GW and age at onset. As not all variables were available for every patient, Fig. 3e presents Model 1 (adjusted only for sex) alongside Model 2, which incorporates each additional adjustment. This approach ensures consistency by analyzing the same subset of patients to account for sample size differences. Notably, no substantial changes in the association between picloram MRS-GW and age at onset were observed after adjustments.
Picloram-induced gene expression alterations are associated with picloram MRS-GW
A high picloram MRS-GW score showed robust associations with EOCRC patients; however, validating whether the picloram MRS reflects actual picloram exposure is not feasible, as data on picloram-related DNA methylation effects are, to our knowledge, unavailable. Alternatively, we validated the pesticide MRSs using changes in gene expression induced by pesticides. Our rationale was that if the gene expression changes induced by pesticide exposure are reflected in the respective MRS, this would support that the MRS represents the molecular effects of pesticide exposure. To this end, we analyzed RNA-seq data (GSE262419) from induced pluripotent stem cell (iPSC)-derived cardiomyocytes exposed to a wide range of chemicals (N = 67) including several pesticides overlapping with those in our study. To identify gene expression alterations for each pesticide, we performed differential gene expression analysis (DGEA) using pesticide exposure as a continuous variable (0.2 µM, 1 µM and 10 µM). Differentially expressed genes (P < 0.005) were then used to generate single-sample gene set enrichment analysis (ssGSEA) scores in TCGA-COAD. The Spearman correlation between each pesticide’s ssGSEA score and its corresponding MRS was tested, identifying notable correlations for dicamba (six genes: R = −0.15; P = 0.025), heptachlor (148 genes: R = −0.15; P = 0.017) and picloram (37 genes: R = 0.44; P = 2 × 10−12) (Fig. 3f), whereas nonsignificant trends were obtained for acetochlor (16 genes: R = −0.015; P = 0.81), atrazine (90 genes; R = −0.07; P = 0.28), lindane (38 genes: R = 0.002, P = 0.98) and metolachlor (51 genes; R = 0.049, P = 0.46). The observed correlations between picloram and dicamba-specific MRSs and ssGSEA scores suggest that these MRSs capture molecular footprints of exposure, supporting their potential as proxies for true pesticide exposure.
Robustness testing for picloram CpGs, patient categorization and the ssGSEA score
To assess robustness, we performed permutation tests to ensure that the observed associations stem from biological relationships rather than artifacts of CpG selection, patient classification or gene selection. CpGs in the picloram MRS-GW ranked 13th in significance among 10,000 permutations (Fig. 3g). Furthermore, patient classification permutation identified age-based classification as the second most significant, based on picloram MRS-GW, among 1,000 permutations (Fig. 3g). CpG site permutations for MDS, education levels, smoking-Maas and obesity are described in Extended Data Fig. 8a and onset categorization permutations in Extended Data Fig. 8b. Finally, permutation testing ranked the picloram-specific ssGSEA score 6th among 1,000 permutations (Fig. 3h), whereas dicamba and heptachlor ranked 71st and 300th, respectively. These results confirm the robustness of CpGs in picloram MRS-GW, patient categorization, and the genes included in the picloram and dicamba ssGSEA scores.
Molecular differences between high and low picloram MRS-GW
Given the robust association between the picloram MRS-GW and younger age at diagnosis, we investigated its relationship with tumor mutations and gene expression. Comparing tumors in the lowest and highest picloram MRS-GW terciles, we identified 75 differentially mutated genes (P < 0.05) in the full dataset, including 42 mutated only in the high exposure (Extended Data Fig. 9). Due to known differences between MSS and MSI, we performed a stratified analysis (Supplementary Table 10). In MSS tumors (Fig. 4a), eight genes were mutated differentially, with one gene restricted to high picloram exposure and five genes restricted to low exposure. Finally, in MSI tumors, 25 genes were mutated differentially (Fig. 4c), including 11 genes unique to low exposure and 10 genes unique to high exposure. The mutation distribution of the identified genes is presented in Fig. 4a–d.
Fig. 4: Molecular differences between high and low picloram MRS-GW.The alternative text for this image may have been generated using AI.
a,b, Tumors were divided into terciles of the picloram MRS-GW, and tumors in the first and third terciles were compared using Fisher’s exact test to identify differentially mutated (P < 0.05) genes. Differentially mutated genes in MSS tumors are shown with the percentage of tumors with a gene mutation in the first or third tercile (a) and their distribution across all MSS tumors, sorted on the picloram MRS (b). c,d, Similarly, the differentially mutated genes in MSI tumors are shown with the percentage of tumors having the gene mutated in the first or third tercile (c) and their distribution across all MSI tumors (d). e, Comparison of gene expression between tumors in the first and third tercile of the picloram MRS-GW using negative binomial generalized linear models. The analysis was adjusted for age, sex, MSI status, tumor purity and tumor location (left/right) and significance thresholds were set at an absolute log2FC > 1.2 and a Padj. < 0.05, resulting in 25 differentially expressed genes with the top 6 genes highlighted. f, NESs for the significantly down- and upregulated pathways in low picloram exposure, using the differential expression results.
DGEA adjusted for age, sex, MSI status, tumor purity and tumor location (left/right) identified 25 differentially expressed genes (Padj. < 0.05, |log2fold-change (FC) | > 1.2), including 16 downregulated genes in low picloram exposure (Fig. 4c and Supplementary Table 11). MSS-stratified analysis identified four differentially expressed genes, whereas MSI-stratified analysis identified 244 genes (Supplementary Table 11). The subsequent GSEA using all samples revealed nine enriched pathways (Padj. < 0.05, |normalized enrichment score (NES)| > 1), including upregulation of the Wnt/β-catenin signaling in low picloram exposure, consistent with APC mutations and downregulation of immune response-related pathways, such as complement and IL6-JAK-STAT3 signaling.
Young tumors associated with picloram exposure
Current CRC classification into early- or late-onset relies on age at diagnosis; however, variability in the interval between tumor initiation and diagnosis makes age at diagnosis an unreliable indicator of the tumor’s actual age26. We therefore evaluated single-base substitution signature 1 (SBS1), a mitotic division-related mutational signature, as a proxy for tumor age. SBS1 reflects the cumulative number of mitotic divisions and captures the replicative history of cells across premalignant and malignant stages27,28. We used the SBS1 score from the Catalogue of Somatic Mutations in Cancer, deconvolved by the Pan-Cancer Analysis of Whole Genomes consortium using established methods with high accuracy (≥0.827)28, and no samples were excluded based on accuracy (Extended Data Fig. 10a). TCGA-COAD patients with DNA methylation and mutational signatures data were included, excluding MSI tumors due to distinct mutational mechanisms (Extended Data Fig. 10b)29,30. SBS1 distribution across age groups early-, intermediate- (IOCRC: aged 50–69 years) and late- (N = 173) onset is detailed in Extended Data Fig. 10c.
Picloram MRS-GW was associated significantly with early-onset versus late-onset cases (N = 25 versus 72; OR: 2.99 [95% CI: 1.70–5.85]; P = 4.27 × 10−4). Next, we employed an SBS1 score threshold, identifying 72 young (SBS1 < 60) and 101 old tumors (SBS1 ≥ 60). The new patient categorization underscored a significant association with picloram MRS-GW (OR: 1.84 [95% CI: 1.31–2.66]; P = 6.57 × 10−4) (Extended Data Fig. 10d). The chronological age distribution for these SBS1-defined groups is shown in Extended Data Fig. 10e. This association remained significant using a non-age-adjusted MRS-GW when adjusting for sex (P = 4.15 × 10−4) and further for age at diagnosis (P = 5.98 × 10−3), confirming independence from chronological age. The SBS1 score was associated strongly with tumor mutation burden (TMB; R = 0.66, P < 2.2 × 10−16). Consequently, adjusting the original model (P = 6.57 × 10−4) for TMB attenuated the association (P = 3.23 × 10−2). To further investigate whether TMB could serve as a marker for tumor age, we performed linear regression models adjusted for sex, showing a stronger association between the picloram MRS-GW and SBS1 score (P = 2.11 × 10−5) than TMB (P = 0.0148), suggesting that although TMB and SBS1 are highly correlated, SBS1 captures the age-specific signal more effectively.
County-level pesticide uses and EOCRC incidence across the USA
Using MRSs as proxies, we identified associations between age at onset and several pesticides. We examined pesticide use intensity and EOCRC incidence rates (IR) across overlapping counties in California, Connecticut, Georgia, Iowa, New Mexico, Utah and Washington in the USA. Pesticide use estimates (1992–2012) were extracted from the Pesticide National Synthesis Project, and the age-adjusted EOCRC IR from the Surveillance, Epidemiology and End Results (SEER) for eight registries (SEER8). Among 225 pesticides with sufficient data (≥10 years and ≥50 total observations), 62 were associated with EOCRC IR in linear mixed models adjusted for years of data collection and county-level random effects (Supplementary Table 12). After additional adjustment for seven county-level socioeconomic status (SES) indicators31, including median household income, median house value, median rent, percent below 150% of the poverty line, education index32, percent working class and percent unemployed, 27 pesticides remained significant (Fig. 5a and Supplementary Table 13).
Fig. 5: Pesticide use intensity is associated with EOCRC incidence.The alternative text for this image may have been generated using AI.
County-level pesticide use intensity estimates (1992–2012) were matched with EOCRC incidence rates. a, Heatmap showing associations between 62 pesticides (y axis) and EOCRC IRs, adjusted for seven SES indicators (x axis). b, The 27 pesticides (y axis) that remained significant were further adjusted for each of the other 26 pesticides (x axis). Pesticides that lost significance in the baseline model without pesticide adjustment are shown in dark pink, whereas those that were significant in the basic model but lost significance after pesticide adjustment are displayed in light pink. Cases with insufficient datapoints for testing are shown in gray. c, The five most robust pesticides were analyzed jointly, adjusting for the remaining pesticides and SES indicators. The left y axis represents the top five pesticides, including atrazine, esfenvalerate, glyphosate, nicosulfuron and picloram, whereas the right y axis reflects for each of them the four pesticides they are adjusted for. The x axis shows the adjustments for only the pesticide or the pesticide and each of the SES indicators. NS, nonsignificant.
As several pesticides were used within counties, we tested robustness by adjusting each of the 27 pesticides for the remaining 26 (Fig. 5b and Supplementary Table 13). Since merging the pesticide data can reduce the number of overlapping datapoints substantially, we first tested baseline models without pesticide adjustment to distinguish true attenuation from reduced data overlap. Reduced overlap led to frequent loss of significance, particularly for fenbuconazole (0.84), propargite (0.80) and flumioxazin (0.65) (Fig. 5b). Among the remaining 24 pesticides, glyphosate and picloram remained significant in 21 of 23 pairwise models, esfenvalerate and nicosulfuron in 15, and atrazine in 12, with others remaining significant in ten or fewer models.
Integrated pesticide- and SES-adjusted analyses of the five most robust pesticides showed limited consistency for atrazine and esfenvalerate, each significant in only one pesticide-adjusted model and moderate robustness for nicosulfuron, which remained significant in two models. Glyphosate showed stronger robustness, maintaining significance in three models, including two with all SES adjustments. Picloram demonstrated the most consistent association, remaining significant in all four pesticide-adjusted models and across most SES-adjusted models (Fig. 5c and Supplementary Table 14). Together with the MRS findings, these results highlight picloram as a potentially important environmental factor contributing to EOCRC risk.