The mouse experiments were approved and performed in accordance with the guidelines of the Institutional Animal Care and Use Committee of the Third Military Medical University (TMMU, Chongqing, China) or Jinfeng Laboratory (Chongqing, China). All the mice were housed in a pathogen-free facility with a 12 h light/dark cycle at the TMMU or Jinfeng laboratory. All the mice were provided food and purified water ad libitum. Human CRC tissue samples were collected in accordance with the guidelines of the Second Affiliated Hospital of Army Medical University (Protocol No. 2019-YANDI 103-01) and the Cancer Hospital of the Chinese Academy of Medical Sciences (Protocol No. KY2023166).
Chemicals
Dimethyl sulfoxide (DMSO; #D8371) and sodium butyrate (NaBu; #156-54-7) were purchased from Solarbio (Beijing, China). DMXAA (#117570-53-3), C-176 (#314054-00-7), Cyclosporin A (CsA; #59865-13-3), mitoquinone mesylate (MitoQ; #845959-50-4), ethidium bromide (EtBr; #1239-45-8), decitabine (#2353-33-5), PEG300 (#25322-68-3), Tween 80 (#9005-65-6), MG132 (#133407-82-6), riboflavin phosphate (sodium), (#HY-B0964), Herbacetin 3,8-O-diglucoside, (HY-N10042), Diosmin (HY-N0178), rutin (HY-N0148), DAPTA (HY-P1034), adenosine 5′-diphosphoribose (sodium) (HY-100973A), Bimosiamose (HY-106139), and hypericin (HY-N0453) were obtained from MedChemExpress (MCE, Shanghai, China). VBIT4 (#2086257-77-2) and N-acetylcysteine (NAC; #616-91-1) were acquired from TargetMol (BOS, MA, USA).
Cell culture
The CRC cell lines MC-38, CT-26, and HCT-116 were maintained in our laboratory.10,45,46 All the cells were authenticated and tested for mycoplasma. Primary mouse macrophages and cell lines were cultured with Dulbecco’s modified Eagle’s medium (DMEM) and high-glucose medium (#C3113, VivaCell, Shanghai, China) supplemented with 10% fetal bovine serum (FBS, #164210-50, Procell, Wuhan, China) at 37 °C in a humidified 5% CO2 atmosphere.
Mice
Wild-type C57BL/6 and BALB/c mice and BALB/c nude mice were obtained from GemPharmatech (Jiangsu, China). C57BL/6 J mice harboring a loxP-flanked (fl) allele of exon 2 of Acads (Acadsfl/-) were obtained from Cyagen Biosciences (Suzhou, Jiangsu, China). Acadsfl/fl mice were crossed with Vil1-Cre mice (Cyagen Biosciences, Suzhou, Jiangsu, China) via a three-step backcrossing procedure to obtain Acads intestinal-conditional knockout mice (Acadsfl/fl-Vil1Cre).
Mouse models of subcutaneous tumors
A mouse model was constructed as described in our previous work.10 Briefly, 100 μL of PBS containing either MC-38 or CT-26 CRC cells (2 × 106) or without cells was subcutaneously injected into the groin of 6-week-old male C57BL/6 or BALB/c mice. Approximately three weeks post-injection, the mice were euthanized, and the subcutaneous tumor nodules were harvested, imaged, and weighed.
Mouse models of peritoneal carcinomatosis
A mouse model was generated as described in our previous studies.45 Briefly, 100 μL of PBS containing MC-38 or CT-26 CRC cells (2 × 106) or without cells was intraperitoneally injected into 6-week-old male C57BL/6 or BALB/c mice. Approximately two weeks post-injection, the mice were euthanized, and the peritoneal tumor nodules were harvested, imaged, and weighed.
Establishment of a colitis-associated cancer model
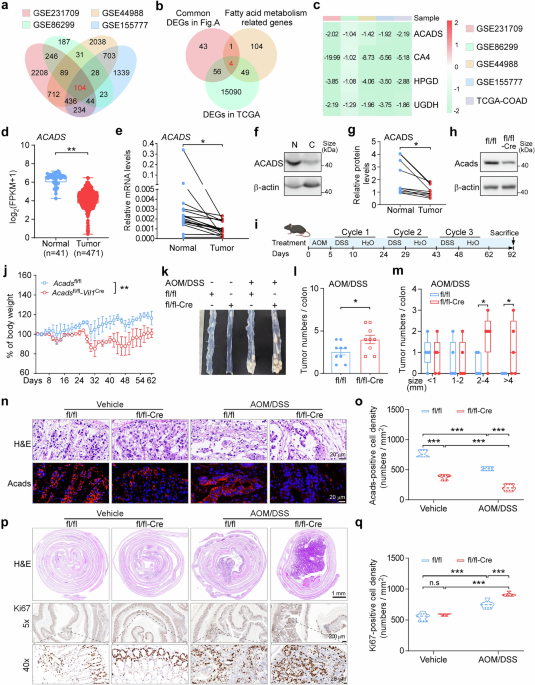
Acadsfl/fl and Acadsfl/fl-Vil1Cre mice (8–12 weeks) were intraperitoneally injected with 10 mg/kg azoxymethane (AOM; #A5486, Sigma). Five days post-injection, the mice underwent three cycles of 2.5% dextran sodium sulfate (DSS; #160110, MP Biomedicals) treatment. Each cycle consisted of administering DSS in their drinking water for 5 consecutive days, followed by a 2-week period of normal water. The body weights of the mice were monitored every two days. After completing the three cycles, the mice were maintained on a regular diet until day 92 post-AOM injection, at which point the colon was excised for analysis.
Generation of Acads-knockdown and Acads-overexpressing MC-38 cells
Lentiviral vectors for Acads knockdown (targeting sequences: #1: 5′-CGCATCACTGAGATCTACGAA-3′; #2: 5′-CCTGGATTGTGCTGTGAAGTA-3′) with puromycin resistance and for the overexpression of C-terminal HA-tagged Acads with hygromycin B resistance were obtained from Genechem (Shanghai, China). MC-38 cells were infected with the respective lentiviruses following the manufacturer’s instructions. After infection, the cells were treated with the corresponding antibiotics for several days until all noninfected cells were eliminated. The successfully transduced cells were then harvested and validated for their Acad expression levels via qPCR and Western blot analysis.
Generation of mito-Dnmt1 knockout and overexpression in MC-38 cells
Mitochondrial Dnmt1-specific knockout MC-38 cells were generated via the pWST-lenti-CRISPR/Cas9 system (Chongqing Western Biomedical Technology Co., Ltd., Chongqing, China). A guide RNA targeting the uORF sequence (5′-CCCCACTCTCTTGCCCTGTG-3′) was designed to disrupt the MLS. The generated CRISPR/Cas9 lentivirus was used to infect MC-38 cells at 50% confluence. Thirty-six hours post-transfection, puromycin was added to the culture medium to select successfully transduced cells. Positive cells were then dissociated and plated into 96-well plates by limiting dilution to obtain single-cell clones. Once colonies formed, genomic DNA was extracted, and the targeted region was amplified via PCR for cloning and subsequent sequencing validation.
For mitochondrial Dnmt1 overexpression, a lentiviral vector containing the full-length Dnmt1 sequence with its uORF encoding the MLS at the N-terminus and a blasticidin resistance marker was packaged by Sangon Biotech (Shanghai, China). MC-38 cells were infected with the mito-Dnmt1 overexpression lentivirus for 48 h, followed by blasticidin selection for several days until all noninfected cells were eliminated. The successfully transduced cells were harvested and validated for mito-Dnmt1 expression via qPCR and Western blot analysis.
Cell viability assay
Acads-knockdown MC-38 cells and their respective negative control cells (5 × 103) were seeded in 96-well plates. After the cells adhered to the bottom of the plates, this time point was designated 0 h. The cell counting kit-8 (CCK-8) working solution (#K009-500, Zetalife, Menlo Park, CA, USA) was added to the plates at predetermined time points (0, 12, 24, 36, and 48 h), followed by an additional incubation for 1 h. Subsequently, the optical density (OD) of each well was measured at 450 nm via a Synergy H1 microplate reader (BioTek, Winooski, VT, USA).
Cell proliferation detection
Acads-knockdown MC-38 cells and their respective negative control cells (5 × 104) were planted in 6-cm petri dishes and cultured in 4 mL of complete medium for 7 days. After the incubation period, the cells were trypsinized and resuspended in an equal volume of PBS. The cell density was subsequently quantified via a hemocytometer.
Apoptosis analysis
Acads-knockdown MC-38 cells and their respective negative control cells (1 × 105) were seeded in 12-well plates. Upon reaching 80% confluence, both the suspended cells in the culture medium and the adherent cells were collected and combined for staining via the Annexin V-FITC/PI Apoptosis Detection Kit (#556547, BD Biosciences, San Jose, CA, USA) according to the manufacturer’s protocol. Apoptosis analysis was then performed via a CytoFLEX flow cytometer (Beckman Coulter, Ltd., Miami, FL, USA). Data were acquired and analyzed via CytExpert software (version 2.4.0.28; Beckman Coulter, Inc.; Brea, CA, USA).
Quantitative real-time PCR (qPCR)
mRNAs from experimental cells were extracted with an Eastep® Super Total RNA Extraction Kit (#LS1040, Promega, Beijing, China). One microgram of total RNA was reverse transcribed into cDNA using PrimeScript™ RT Master Mix (#RR036, TaKaRa, Beijing, China). qPCR was performed in a 20 µL reaction volume containing TB Green premix (#RR820a, TaKaRa, Beijing, China) via the CFX Connect™ Optics Module system (Bio-Rad, Hercules, CA, USA) following the instructions of the kit. Relative gene expression levels were calculated via the 2−∆∆CT method, with β-actin serving as the housekeeping gene for normalization. The primer sequences are provided in Supplementary Tables 1 and 2.
Western blotting
The CRC cell lines MC-38 or CT-26 were plated in 6-well plates. Upon reaching 70%-80% confluence, the cells were washed with precooled PBS and lysed on ice in cell lysis/IP buffer (#P0013, Beyotime, Shanghai, China) supplemented with protease inhibitors (#HY-K0011, MedChemExpress, Shanghai, China). The protein concentration of the lysates was detected via an enhanced BCA protein assay kit (#P0010, Beyotime, Shanghai, China). All samples were adjusted to an equal concentration and denatured with 5× loading buffer at 100 °C for 10 min. Equal amounts of protein from each sample were then loaded into an 8% or 10% sodium dodecyl sulfate‒polyacrylamide gel (SDS‒PAGE) for electrophoresis. The resolved proteins were transferred onto PVDF membranes, which were subsequently blocked with 5% nonfat milk in Tris-buffered saline containing 0.1% Tween 20 (TBST) for 1 h at room temperature. The membranes were washed with TBST and incubated overnight at 4 °C with primary antibodies, including ACADS (#AP75024, Abcepta, Jiangsu, China), DNMT1 (#NB100-56519AF594, NOVUS), cGAS (Cat#31659, Cell Signaling Technology), p-STING (#PA5-105674, Invitrogen), STING (#A3575, ABclonal), p-TBK1 (#AP1026, ABclonal), TBK1 (#AF8103, Beyotime), p-IRF3 (#4947, Cell Signaling Technology), IRF3 (#4302, Cell Signaling Technology), BAX (#A0207, ABclonal), VDAC1 (#A19707, ABclonal), VDAC3 (#55260-1-AP, Proteintech), HSP60 (#AF0186, Beyotime), PHB1 (#10787-1-AP, Proteintech), Lamin B1 (#ab16048, Abcam), HA (#51064-2-AP, Proteintech), TFAM (#A13552, ABclonal), NRF1 (#12482-1-AP, Proteintech), PGC1α (#66369-1-Ig, Proteintech), Twinkle (#18793-1-AP, Proteintech), POLG (#AP14948B, Abcepta), and β-actin (#66009-1-Ig, Proteintech). Following incubation with primary antibody, the membranes were washed 3 times with TBST and then hybridized with the corresponding secondary antibody at room temperature for 2 h. In certain instances, the membrane was subjected to washing with primary and secondary antibody removal solution (#P0025, Beyotime), followed by reincubation with either the indicated antibody or a loading control β-actin antibody, along with the corresponding secondary antibodies. The chemiluminescence signals of the target proteins were collected via a chemiluminescence instrument after the addition of the enhanced chemiluminescence substrate. Protein band density was quantified via ImageJ software (version 1.54g; https://imagej.net/ij/).
Cytoplasmic mtDNA analysis
Experimental cells (5 × 105) were seeded in 6-well plates. When the cells reached 70%-80% confluence, they were trypsinized, collected by centrifugation at 400 × g for 5 min, and then resuspended in PBS. The cell suspension was equally divided into two aliquots. The first aliquot was lysed on ice using 1% IGEPAL CA-630 (#ST2045, Beyotime, Shanghai, China) for 15 min, followed by centrifugation at 16,000 × g for 15 min at 4 °C. The resulting supernatant, representing the cytoplasmic fraction, was transferred to a new tube for subsequent genomic DNA extraction via a commercial kit (TIANGEN, #DP304-03). The second aliquot was processed for total genomic DNA extraction following the manufacturer’s protocol. To quantify the mtDNA content in the cytoplasmic fraction, qPCR was performed using primers specific for the mitochondrial ND1 and D-loop, while the nuclear Tert gene from total genomic DNA served as an internal reference for normalization. The primer sequences are shown in Supplementary Table 1.
Mitochondrial ROS detection
Mitochondrial ROS levels were assessed via the MitoSOX Red mitochondrial superoxide indicator (#1003197-00-9; MedChemExpress, Shanghai, China) according to the manufacturer’s protocol. In brief, experimental cells (1 × 105) were seeded in 12-well plates and allowed to reach 70–80% confluence. The cells were then trypsinized, harvested by centrifugation at 600 × g for 5 min, and resuspended in 1 mL of culture medium containing 100 nM MitoSOX Red working solution. Following a 30 min incubation at room temperature in the dark, the cells were centrifuged at 400 × g for 5 min and washed twice with PBS. The cell suspension was subsequently filtered through a 200-mesh filter and subjected to flow cytometry analysis via appropriate excitation/emission settings (510/580 nm) for MitoSOX Red detection. The fluorescence intensity was quantified via CytExpert software (version 2.4.0.28; Beckman Coulter, Inc.; Brea, CA, USA).
Mitochondrial membrane potential (MMP) assay
The MMP was measured via the fluorescent probe tetramethylrhodamine methyl ester (TMRM) (#115532-49-5; MedChemExpress, Shanghai, China) following the manufacturer’s protocol. Briefly, experimental cells were seeded in 12-well plates and cultured until 80% confluence. The cells were then trypsinized and collected by centrifugation at 500 × g for 5 min at room temperature. The cell pellet was resuspended in 1 mL of serum-free medium containing 20 μM TMRM working solution and incubated at 37 °C in a 5% CO2 atmosphere for 30 min. After incubation, the cells were washed with PBS following centrifugation at 500 × g for 5 min and finally resuspended in 200 μL of PBS. Flow cytometry analysis was performed using a 488 nm excitation laser with fluorescence emission collected at 570 nm (±10 nm) through the appropriate bandpass filter. Data acquisition was performed via a CytoFLEX flow cytometer (Beckman Coulter, Ltd., Miami, FL, USA), whereas data analysis was conducted via CytExpert software (version 2.4.0.28, Beckman Coulter, Inc., Brea, CA, USA).
ATP content assay
The intracellular ATP levels were quantified via an enhanced ATP assay kit (#S0027; Beyotime, Shanghai, China) following the manufacturer’s instructions. In brief, experimental cells cultured in 6-well plates were washed twice with PBS upon reaching 80% confluence, followed by the addition of lysis buffer. The cell lysates were collected via centrifugation at 12,000 × g for 5 min at 4 °C. The resulting supernatants were then mixed with the ATP detection working solution at a 1:1 ratio (v/v). Luminescence intensity was measured via a multimode microplate reader (SpectraMax iD5, Molecular Devices, USA). ATP concentrations were determined via interpolation from a standard curve generated with known ATP concentrations (0.1–10 μM), and the values were normalized to the total protein content determined via a BCA assay.
Mitochondrial calcium level analysis
The experimental cells were seeded in 6-well plates and cultured until they reached 80% confluence. The cells were then trypsinized and harvested via centrifugation at 500 × g for 5 min at room temperature. Following washing with PBS, the cell pellets were resuspended in PBS containing 10 μM Rhod-2 AM (#145037-81-6, MedChemExpress, Shanghai, China) and incubated at room temperature for 30 min in the dark. After incubation, the cells were collected by centrifugation at 500 × g for 5 min and washed twice with PBS. Finally, the cells were resuspended in PBS for fluorescence intensity analysis via flow cytometry. The data were further analyzed via FlowJo (v10.8.1).
Liquid chromatography‒tandem mass spectrometry (LC‒MS/MS) and coimmunoprecipitation assay
LC‒MS/MS and coimmunoprecipitation assays were performed following the protocol established in our previous study.47 In brief, 4 μg of specific antibodies against Acads (#16623-1-AP, Proteintech) or Dnmt1 (#NB100-56519AF594, Novus Biologicals) and their respective IgGs were preincubated with magnetic beads (Sera-Mag SpeedBead Protein A/G, 17152104010150, GE) from a 40 μL suspension at 4 °C overnight. After incubation, the bead‒antibody complex was washed three times with cell lysis/IP buffer (#P0013, Beyotime, Shanghai, China). Subsequently, 500 μg of the protein lysate from MC-38 cells was added to the mixture, which was subsequently incubated overnight at 4 °C with gentle rotation. The protein bound to the beads was then eluted via 2× protein loading buffer at 100 °C for 10 min, and the eluate was subjected to SDS‒PAGE and stained with Coomassie brilliant blue G250 (#P0003S, Beyotime, Shanghai, China). The gels containing all the proteins from the IgG, Acads, and Dnmt1 groups were subsequently sent to Shanghai Applied Protein Technology Co., Ltd. (Shanghai, China) for LC‒MS/MS analysis. Additionally, the eluate was further separated and analyzed by Western blotting using antibodies against Acads (#16623-1-AP, Proteintech), Dnmt1 (#NB100-56519AF594, Novus Biologicals), or ubiquitin (#3933, Cell Signaling Technology) to perform the coimmunoprecipitation assay.
Protein expression, purification, and in vitro protein binding assay
The coding sequences of Acads and Dnmt1 were cloned and inserted into the glutathione S-transferase (GST)-containing pGEX-4T-3 plasmid. The recombinant plasmids, along with the empty vector, were transformed into E. coli BL21 cells. The transformed cells were cultured in 2× YT medium at 37 °C with shaking until they reached the logarithmic growth phase (OD₆₀₀ = 0.4–0.6). Protein expression was induced by adding isopropyl β-D-1-thiogalactopyranoside (IPTG) to a final concentration of 1 μM, followed by incubation at 20 °C for 8 h with shaking at 200 rpm. After induction, the bacterial cells were resuspended in PBS supplemented with protease inhibitor cocktails and subjected to ultrasonic lysis on ice to disrupt the cell membranes. The cell lysates were collected via centrifugation at 12,000 × g for 5 min at 4 °C. The supernatant was then incubated with glutathione Sepharose 4B beads (#17-0756-01, GE Healthcare, Sweden) at 4 °C overnight under gentle rotation. Following incubation, the supernatant was discarded, and the agarose beads were thoroughly washed five times with cold PBS to remove nonspecifically bound proteins. For subsequent analysis, the washed agarose beads were used for a second round of immunoprecipitation. The eluate was incubated overnight at 4 °C with gentle rotation using lysates from HEK293T cells transfected with either a Dnmt1-Flag plasmid or an empty control plasmid. After incubation, the beads were collected and washed five times with cold lysis buffer. Finally, the beads were eluted with glutathione elution buffer and mixed with 2× loading buffer for Western blot detection.
Immunofluorescence and superresolution imaging
The immunofluorescence assay was performed following the protocol established in our previous study.47 Briefly, MC-38 cells were seeded onto glass coverslips placed in 12-well plates. Upon reaching 70% confluence, the cells were gently rinsed with PBS and fixed with 4% paraformaldehyde at room temperature for 15 min. Following fixation, the cells were treated with 5% bovine serum albumin (BSA) prepared in 0.3% Triton X-100 solution to block nonspecific antigen binding sites, and this blocking step was carried out at 37 °C for 30 min. The cells were then incubated overnight at 4 °C with primary antibodies targeting Hsp60 (#A85438, Antibodies), Acads (#16623-1-AP, Proteintech), Dnmt1 (#NB100-56519AF594, Novus Biologicals), 5-Methylcytosine (5-mC; #39649, Proteintech), or DNA (#CBL186, Merck Millipore). After thorough washing with PBS, the cells were exposed to a fluorophore-conjugated secondary antibody for 2 h at room temperature in a dark and humidified chamber. Nuclei were counterstained with Antifade Mounting Medium with DAPI (#P0131, Beyotime, Shanghai, China) in the dark for 20 min. Finally, the stained samples were imaged and captured via either a laser-scanning confocal microscope (IXplore SpinSR, Olympus) or a Multi SIM X superresolution microscope (NanoInsights, Beijing, China) for superresolution imaging.
Proximity ligation assay (PLA)
MC-38 cells were seeded onto glass coverslips placed in 12-well plates. Upon reaching 70% confluence, the cells were gently rinsed with PBS and fixed with 4% paraformaldehyde at room temperature for 15 min. After fixation, the cells were washed three times with PBS, and the PLA was performed following the protocol provided in the Duolink In Situ Red Starter Kit Mouse/Rabbit Kit (#DUO92101, Sigma‒Aldrich). Next, a blocking solution was applied to prevent nonspecific binding of the cellular antigens by incubation at 37 °C for 1 h. Antibodies against Acads (#16623-1-AP, Proteintech) and Dnmt1 (#NB100-56519AF594, Novus Biologicals), either individually or in combination, were prepared in Probemaker PLA Probe Diluent and incubated with the cells overnight at 4 °C. Next, the PLA probes were ligated to their respective antibodies and amplified in the appropriate buffer at 37 °C for 1 h. The nuclei were then stained with Duolink In Situ Mounting Medium containing DAPI and incubated for 20 min at room temperature in the dark. Finally, images were captured via a confocal microscope (IXplore SpinSR, Olympus).
Mitochondrial isolation
Mitochondria were isolated via a mitochondria isolation kit (#89874; Thermo Fisher Scientific) following the manufacturer’s protocol. Briefly, experimental cells were cultured in 15-cm petri dishes. Upon reaching 80% confluence, the cells were gently washed with PBS, trypsinized, and collected by centrifugation at 850 × g for 2 min. The cell pellet was resuspended in ice-cold Reagent A, followed by the addition of other isolation reagents and thorough mixing. The lysate was then centrifuged at 700 × g for 10 min to remove nuclei and cellular debris. The supernatant, containing mitochondria, was further centrifuged at 3000 × g for 15 min to pellet the mitochondria. The resulting pellet represented the mitochondrial fraction, while the supernatant contained the cytoplasmic components.
mtDNA methylation analysis
The analysis of mtDNA methylation was performed via the MethylFlashTM Global DNA Methylation (5-mC) ELISA Easy Kit (Colorimetric) (#P-1030, EpigenTek), which strictly adhered to the manufacturer’s protocol. In brief, mtDNA was extracted from isolated mitochondria via a genomic DNA extraction kit (TIANGEN, #DP304-03). An equal amount of mtDNA (100 ng) was diluted with binding solution and then transferred to strip wells with high DNA affinity, followed by incubation at 37 °C for 1 h. Subsequently, each well was washed with washing buffer, and a 5-mC detection complex solution was added and incubated at room temperature for 50 min. After another wash with diluted washing buffer, the developer solution was added to each well. Following a 4 min reaction, the stop solution was added to halt the enzymatic reaction, and the absorbance was measured at 450 nm via a microplate reader (SpectraMax iD5, Molecular Devices, USA). The percentage of 5-mC was determined on the basis of a standard curve.
Hematoxylin and eosin (H&E) staining
H&E staining of the tissues was performed following the protocol established in our previous study.10 Briefly, the colorectum of each mouse was rinsed with PBS and prepared via the Swiss-roll technique. The tissue was then fixed in 4% paraformaldehyde and embedded in paraffin. The paraffin-embedded blocks were subsequently sectioned into slices, which underwent a series of processing steps, including dewaxing, dehydration with graded ethanol solutions, and rehydration with distilled water. Following pretreatment with a prestaining solution, the tissue sections were stained with H&E solution (#C0105, Beyotime, Beijing, China). After staining, the slides were dehydrated and mounted with a coverslip. The stained slides were finally examined and imaged via a light microscope.
Immunohistochemistry and tyramide signal amplification (TSA)-based multiple immunohistochemistry
The rehydrated slides were subjected to high-pressure-induced epitope retrieval in citrate buffer (pH 6.0). After cooling to room temperature, the slides were washed three times with PBS and treated with 3% H2O2 in the dark for 25 min to block endogenous peroxidase activity. The slides were subsequently incubated with 10% rabbit serum or 3% BSA at room temperature for 30 min to block nonspecific antigen binding sites, followed by overnight incubation with the primary antibody at 4 °C in a humidified chamber. After being washed three times with PBS, the slides were incubated with an HRP-labeled secondary antibody for 1 h at room temperature. For conventional immunohistochemistry, the slides were washed three times with PBS and treated with DAB substrate to develop the chromogenic reaction. For triple or quadruple immunohistochemistry, the slides were incubated with fluorophore-labeled TSA at room temperature for 10 min. After incubation, the slides were washed three times with TBST and subjected to microwave-induced epitope retrieval in citrate buffer. This was followed by two or three additional cycles, each comprising blocking nonspecific binding sites, incubation with primary and secondary antibodies, addition of TSA with a distinct fluorophore, and microwave-induced epitope retrieval to stain two or three additional target proteins. Upon completion of the staining process, the slides were treated with DAPI to stain the nuclei, followed by the application of an autofluorescence quencher and mounting with a coverslip. Finally, the results were visualized and captured via a fluorescence microscope (CLIPSE C1, Nikon) and a slide scanner (Pannoramic MIDI, 3DHISTECH). The antibodies used for immunohistochemistry were as follows: Ki67 (#GB11612, Servicebio), STING (#GB111415, Servicebio), GR1 (#GB11229, Servicebio), CD3 (#GB12014, Servicebio), F4/80 (#GB113373, Servicebio), FOXP3 (#GB112325, Servicebio), LY6C (#GB115601, Servicebio), LY6G (#GB11224, Servicebio), CD25 (#GB11612, Servicebio), ACADS (#TA800429S, Thermo Fisher Scientific), DNMT1 (#GB11328, Servicebio), and HSP60 (#GB15243, Servicebio). T-cell populations were defined on the basis of specific surface marker profiles: total T cells as CD3+; CD4+ T cells as CD3+CD8-CD4+; and CD8+ T cells as CD3+ CD4-CD8+. Macrophage populations were categorized as follows: total macrophages, F4/80+; M1-like cells, F4/80+CD206-iNOS+; and M2-like cells, F4/80+iNOS-CD206+. MDSCs were classified as total MDSCs as Gr1+; polymorphonuclear MDSCs (PMN-MDSCs) as CD11b+LYG+LY6C-; and monocytic MDSCs (M-MDSCs) as CD11b+LYG-LY6C+. Regulatory T cells (Tregs) were identified as CD3+CD25+FOXP3+ cells. In addition, the positive cell density of different immune cells and the percentage of colocalized positive cells within tumors were analyzed via the HALO software platform (Indica Labs, USA).
Fluorescence-activated cell sorting (FACS) of human CRC, mouse tumor, and adipose tissues
The human CRC samples and subcutaneous tumors from the mice were minced into 1 mm³ pieces and then digested in 20 mL of RPMI-1640 medium or DMEM supplemented with 1% FBS, 1 mg/mL collagenase IV (#C5138, Sigma), 0.1 mg/mL hyaluronidase (#BS171-1 g, Biosharp Life Science), or 0.1 mg/mL DNase I (#10104159001, Merck Millipore) at 37 °C for 45 min. For adipose tissue processing, epididymal fat pads were harvested, minced, and digested in 10 mL of DMEM containing 1% FBS and 1 mg/mL collagenase IV (#C5138, Sigma) at 37 °C for 45 min. Following complete digestion, the cell suspensions were filtered through a 70 μm strainer and centrifuged at 500 × g for 10 min. The pelleted cells were resuspended in 40% Percoll solution (#17089109, Cytiva) and carefully layered onto 70% Percoll solution, followed by centrifugation at 800 × g for 20 min. The intermediate layer containing the target cells was collected and treated with red blood cell lysis buffer (R1010, Solarbio, China) to remove erythrocytes. After lysis, the cells were centrifuged at 500 × g for 5 min, resuspended in PBS, and incubated with fluorochrome-conjugated antibodies targeting cell surface markers for 45 min at 4 °C. The cells were then washed twice with PBS at room temperature, subsequently resuspended, and filtered through a cell strainer. FACS analysis was then performed using a FACSVerse C6 flow cytometer (BD LSRFortessa). For intracellular marker staining, the cells were fixed and permeabilized before antibody incubation. The antibodies used for FACS analysis were as follows: PerCP/Cy5.5 anti-mouse CD45 (#103132, BioLegend), FITC anti-mouse CD90.2 antibody (#105338, BioLegend), APC/Cy7 anti-mouse F4/80 antibody (#123118, BioLegend), PE anti-mouse CD11c isotype control antibody (#117308, BioLegend), and APC anti-mouse CD206 isotype control antibody (#141708, BioLegend); APC anti-mouse CD11b antibody (#101212, BioLegend), Percp5.5 anti-mouse Gr1 antibody (#108428, BioLegend), and FITC anti-mouse LY6C antibody (#128006, BioLegend); Alexa Fluor 700 anti-mouse CD45 antibody (#103128, BioLegend), PE anti-mouse CD3 antibody (# 100206, BioLegend), APC/Cy7 anti-mouse CD4 antibody (#116020, BioLegend), Percp5.5 anti-mouse CD8a (#100734, BioLegend), and APC anti-mouse IFNγ (#505810, BioLegend); and APC anti-mouse CD25 antibody (#102012, BioLegend), FITC anti-mouse Foxp3-AF488 antibody (#126406, BioLegend). Brilliant Violet 421™ anti-human CD45 antibody (#368521, Biolegend), FITC anti-human CD11B antibody (301329, Biolegend), APC anti-human CD68 antibody (#333813, Biolegend), Brilliant Violet 510™ anti-human CD3 antibody (#317331, Biolegend), PE/Cyanine7 anti-human IFN-γ antibody (#502527, Biolegend), APC/Fire™ 810 anti-human CD25 antibody (#356149, Biolegend), and PerCP-Cy5.5 anti-human FOXP3 antibody (#320212, Biolegend) were used. T-cell populations were identified via the following surface marker profiles: total T cells, CD45+CD3+; CD4+ T cells, CD45+CD3+CD8-CD4+; and CD8+ T cells, CD45 + CD3 + CD4-CD8+. Macrophage populations were characterized as follows: total macrophages, CD90.2-CD45+F4/80+; M1-like cells, CD90.2-CD45+F4/80+CD206-CD11c+; and M2-like cells, CD90.2-CD45+F4/80+CD11c-CD206+. The populations of MDSCs were classified as total MDSCs as CD11b+Gr1+; PMN-MDSCs as CD11b+LYG+LY6C-; and M-MDSCs as CD11b+LYG-LY6C+. Treg cells were identified as CD45+CD3+CD4+CD25+FOXP3+ cells.
RNA sequencing and DNA methylation sequencing
The experimental cells were seeded into 6-well plates and cultured until they reached 80% confluence. Next, the cells were gently washed twice with ice-cold PBS. Subsequently, 500 µL of TRNzol Universal reagent (#DP424, TIANGEN, Beijing, China) was added to lyse the cells. The resulting cell lysates were then collected, immediately placed on dry ice, and promptly transported to TSINGKE (Beijing, China) for transcriptome sequencing. Concurrently, MC-38 cells with Acads knockdown or mito‑Dnmt1 overexpression, along with their corresponding controls, were harvested. Genomic DNA was then extracted in parallel from two sources: (1) whole‑cell genomic DNA was sent to the BGI Center (Shenzhen, Guangdong, China) and TSINGKE (Beijing, China) for DNA methylome sequencing and BSP, respectively; and (2) mtDNA isolated from purified mitochondria was sent to TSINGKE (Beijing, China) for methylation analysis via BSP and third‑generation nanopore sequencing.
Transcriptomic data analysis and visualization
Transcriptomic sequencing data from four distinct datasets (GSE231709, GSE86299, GSE44988, and GSE155777) pertaining to AOM/DSS-induced mouse spontaneous colon cancer models were retrieved from the GEO database. These datasets were subsequently analyzed via the GEO2R online tool to identify differentially expressed genes (DEGs) by comparing the treatment groups to their respective controls. DEGs were defined as those with an absolute log2-fold change (|log2FC|) > 1 and an adjusted p-value < 0.05. Next, an online intersection analysis (http://bioinformatics.psb.ugent.be/webtools/Venn/) was carried out to compare these DEGs with those identified from The Cancer Genome Atlas (TCGA)-Colon Adenocarcinoma (COAD) database, the latter of which were processed via the DESeq2 package (version 1.40.2) in R software (version 4.4.1). Additionally, genes associated with fatty acid metabolism from the HALLMARK_FATTY_ACID_METABOLISM gene set in the MsigDB database were included in the subsequent analysis. The common DEGs identified through this process were then mapped, and their expression changes were visualized via a pheatmap generated via the heatmap package (version 1.0.12) in R software (version 4.4.1).
Single-cell RNA-Seq data analysis and visualization
Single-cell RNA sequencing data (GSE178341)48 from human CRC patient samples were retrieved from the GEO database. The data were subjected to quality control, standardization, and dimensionality reduction clustering analysis via the Seurat package (version 5.1.0) in R software. Tumor cells were subsequently isolated to assess ACADS expression levels. All analyses were performed via default parameters unless otherwise specified. Cell clustering was achieved via the ‘FindClusters’ function in Seurat, with cell types and subtypes identified through nonlinear dimensional reduction (t-SNE) and annotated on the basis of established cell type-specific markers.48,49 From the dataset, we identified 10 pairs of CRC tissue samples with distinct ACADS expression profiles in tumor cells. The low ACADS expression group comprised the following samples: C103_T_1_1_0_c1_v2, C114_T_1_1_0_c1_v2, C119_T_0_2_0_c1_v2, C125_T_1_1_0_c1_v2, C134_T_0_3_0_c2_v2, C136_T_1_1_0_c1_v2, C145_T_1_1_0_c1_v2, C159_T_1_1_0_c1_v3, C165_T_0_0_0_c1_v3, and C166_T_0_0_0_c1_v3. The high ACADS expression groups included C106_T_1_1_0_c1_v2, C122_T_1_1_0_c1_v2, C130_TA_1_1_0_c1_v2, C130_TB_1_1_0_c1_v2, C132_T_1_1_0_c1_v2, C137_T_1_1_0_c1_v2, C139_T_1_1_0_c1_v2, C139_T_1_1_0_c2_v2, C146_T_1_1_0_c1_v2, and C164_T_1_1_0_c1_v3. These 10 sample pairs were subsequently integrated and analyzed via the Seurat package, with batch effects corrected via the Harmony package (version 1.2.1). Following confirmation of differential ACADS expression in tumor cells between the groups, we examined changes in cell proportions within each group on the basis of the established clustering and grouping criteria, followed by statistical analysis.
Immune infiltration and immune epigenetic scoring analysis
Immune infiltration analysis was conducted with the xCell tool (version 1.1.0) in R software according to the expression data of the ACADS, DNMT1, and STING signaling pathways from the TCGA-COAD cohort. The genes associated with immune infiltration patterns were analyzed via multiple algorithms, including CIBERSORT, TIDE, EPIC, TIMER, and XCELL, via the TIMER2.0 online platform (http://timer.cistrome.org/). Additionally, the immune epigenetic score of ACADS patients was evaluated by utilizing the IPS of COAD patients who underwent anti-PD-1/CTLA-4 immunotherapy, as provided in the TCIA database (https://tcia.at/home).
Analysis of tissue microarrays
CRC and adjacent paracancerous tissues from the Cancer Hospital of the Chinese Academy of Medical Sciences (Protocol No. KY2023166) were processed into tissue microarrays and immunohistochemically stained with antibodies against ACADS (#TA800429S, Thermo Fisher Scientific), DNMT1 (#GB11328, Servicebio), and HSP60 (#GB15243, Servicebio). The staining results were analyzed via the HALO software platform (Indica Labs, USA) to quantify key parameters, including positive cell counts, positive areas, and colocalization. These metrics were subsequently used to calculate the ratio of positive cells to the positive area. Additionally, the H score for ACADS expression across different stages of CRC tissues was determined via the following formula: H score = ∑(pi × i) = (percentage of weakly stained cells × 1) + (percentage of moderately stained cells × 2) + (percentage of strongly stained cells × 3). Here, i represents the intensity grading of the positive area: negative (no staining, scored as 0), weakly positive (light yellow, scored as 1), moderately positive (brownish yellow, scored as 2), and strongly positive (dark brown, scored as 3). The term pi denotes the percentage of the positive area corresponding to each intensity grade.
Structure-based virtual screening
Virtual screening for Acads-specific binders was performed by MedChemExpress (Shanghai, China). Briefly, the 3D structure of Acads (PDB ID: 2VIG) was retrieved from the Protein Data Bank, followed by structure optimization, binding site determination, and grid construction. Moreover, 2D structures of small molecules from the MedChemExpress library were converted into 3D conformations. Then, molecular docking was performed on the basis of the geometric and energetic complementarity. The top 15% of the compounds identified in the primary screening were subjected to secondary screening in standard-precision mode. The top 15% of hits from this secondary screening were subsequently further evaluated in a tertiary screening via the extraprecision mode. Finally, the binding interactions and structural stability between Acads and the candidate compounds were manually validated.
Surface plasmon resonance (SPR)
The CM5 sensor chip (Cytiva, BR-1005-30) was immobilized with recombinant ACADS protein (amino acids 25–412) according to standard amine coupling protocols. The binding kinetics between ACADS and hypericin were evaluated via SPR in manual injection mode. A serial dilution of hypericin was prepared for kinetic analysis. The running buffer was delivered at a flow rate of 30 μL/min, with both the association and dissociation phases lasting 150 s each. Multicycle kinetic analysis was conducted, and sensorgrams were plotted with time (seconds) on the x-axis and response units (RUs) on the y-axis. Kinetic parameters were calculated via BIAcore Insight evaluation software (Cytiva, Marlborough, MA, UK) via the steady state affinity model, yielding the equilibrium dissociation constant (KD, M), association rate constant (Ka, 1/Ms), and dissociation rate constant (Kd, 1/s).
Statistical analysis
Statistical analyses were conducted via GraphPad Prism 8. The data from the animal experiments and clinical studies are presented as the means ± SEMs, whereas the other results are expressed as the means ± SDs. Comparisons between two groups were performed via Student’s t tests, and comparisons among multiple groups were analyzed via one-way analysis of variance (ANOVA). Survival curves were evaluated via the log-rank test. The relationship between ACADS and mito-DNMT1 in terms of mean fluorescence intensity (MIF) scores was assessed via Pearson’s correlation analysis. All the experiments were performed with biological replicates and were repeated at least three times. The mice were randomly assigned to experimental groups. A p-value of <0.05 was considered statistically significant.